
Rice Science ›› 2019, Vol. 26 ›› Issue (2): 88-97.DOI: 10.1016/j.rsci.2019.01.002
• Research Papers • Previous Articles Next Articles
Songmei Liu1,2, Jie Jiang1,2, Yang Liu1,2, Jun Meng3, Shouling Xu1, Yuanyuan Tan1, Youfa Li4, Qingyao Shu1,2, Jianzhong Huang1( )
)
Received:2018-09-27
Accepted:2018-11-26
Online:2019-03-04
Published:2018-12-18
Songmei Liu, Jie Jiang, Yang Liu, Jun Meng, Shouling Xu, Yuanyuan Tan, Youfa Li, Qingyao Shu, Jianzhong Huang. Characterization and Evaluation of OsLCT1 and OsNramp5 Mutants Generated Through CRISPR/Cas9-Mediated Mutagenesis for Breeding Low Cd Rice[J]. Rice Science, 2019, 26(2): 88-97.
Add to citation manager EndNote|Ris|BibTeX
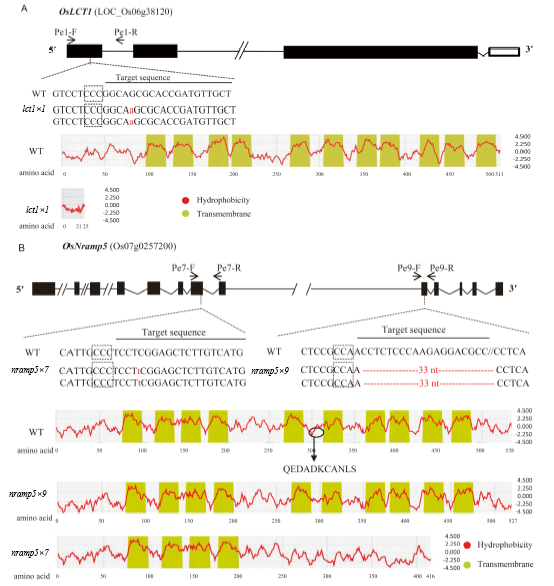
Fig. 1. Schematic diagram of the genes and sgRNA target sites for CRISPR/Cas9-mediated mutagenesis of OsLCT1 (A) and OsNramp5 (B), together with a hydrophobic plot of the wild type (WT) and mutant proteins.Exons, introns, and untranslated regions are indicated by solid boxes, lines, and hollow boxes, respectively. The protospacer adjacent motif sequences (Meharg et al, 2013) prior to the target sequences are boxed with dot lines. The primers Pe1-F/-R, Pe7-F/-R and Pe9-F/-R used for genotyping the mutations and their positions are indicated by arrows. Mutations within or around the target sequences of the mutant lines lct1×1, nramp5×7, and nramp5×9 are shown in lowercase letters (for insertion) or ‘-’ (for deletion). The hydrophobicity (red curves) and the estimated transmembrane domains (filled boxes) of the predicted proteins were drawn using the SOSUI program (http://harrier.nagahama-i-bio.ac.jp/sosui/). For nramp5×9, the position of the deletion is shown in a circle in the WT hydrophobic plot and the deleted amino acid sequence is shown by uppercase letters.
| Plasmid | No. of transgenic plants | No. of plants tested for mutation | No. of mutated plants | |||
| Total | Monoallelic heterozygous | Biallelic heterozygous | Biallelic homozygous | |||
| pH-lct1×1 | 69 | 69 | 15 | 10 | 5 | 0 |
| pH-nramp5×7 | 6 | 6 | 1 | 0 | 1 | 0 |
| pH-nramp5×9 | 49 | 4 a | 4 | 0 | 3 | 1 |
| a Due to technical reasons, we failed to amplify the fragment encompassing the target site in exon 9 ( | ||||||
Table 1. Development and analysis of T0 rice plants for targeted mutagenesis of OsLCT1 and OsNramp5.
| Plasmid | No. of transgenic plants | No. of plants tested for mutation | No. of mutated plants | |||
| Total | Monoallelic heterozygous | Biallelic heterozygous | Biallelic homozygous | |||
| pH-lct1×1 | 69 | 69 | 15 | 10 | 5 | 0 |
| pH-nramp5×7 | 6 | 6 | 1 | 0 | 1 | 0 |
| pH-nramp5×9 | 49 | 4 a | 4 | 0 | 3 | 1 |
| a Due to technical reasons, we failed to amplify the fragment encompassing the target site in exon 9 ( | ||||||

Fig. 2. Mutations identified within and around the target sites of OsLCT1 and OsNramp5 generated through CRISPR/Cas9-mediated genome editing in rice in T1 generation.The protospacer adjacent motif sequences are boxed. The 20-nt target sequences are underlined. Dashes indicate deletions and lowercase letters indicate inserted nucleotides.
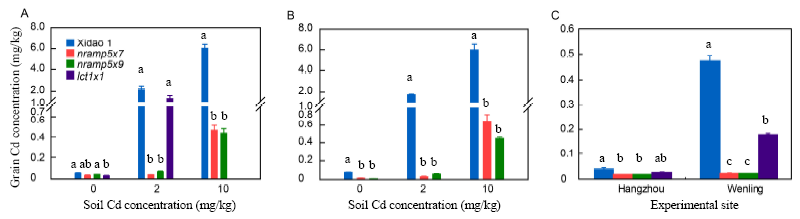
Fig. 3. Cadmium content in brown rice grains of the mutant lines nramp5×7, nramp5×9, lct1×1 and wild type Xidao 1.A, Pot experiment in 2016. B, Pot experiment in 2017. C, Field experiment in 2017. Plants were grown in soil spiked with 2 or 10 mg/kg Cd. Different letters above columns indicate significant difference at the 0.05 level. The data presented are Mean ± SE (n = 3).
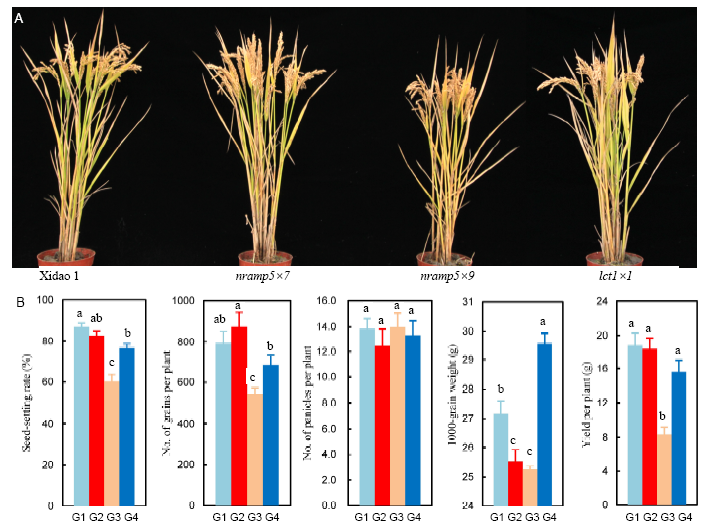
Fig. 4. Plant morphology (A) and yield-related traits (B) of Xidao 1 and its OsLCT1 and OsNramp5 mutants grown in a paddy field at Hangzhou, China, in 2017.G1, Xidao 1; G2, nramp5×7; G3, nramp5×9; G4, lct1×1. Data are presented as Mean ± SE (n = 6). Different letters above columns indicate significant differences at the 0.05 level.
| [1] | Abe T, Ito M, Takahashi R, Honma T, Sekiya N, Shirao K, Kuramata M, Murakami M, Ishikawa S.2017. Breeding of a practical rice line ‘TJTT8’ for phytoextraction of cadmium contamination in paddy fields.Soil Sci Plant Nutr, 63(4): 388-395. |
| [2] | Arao T, Ae N.2003. Genotypic variations in cadmium levels of rice grain.Soil Sci Plant Nutr, 49(4): 473-479. |
| [3] | Arao T, Ishikawa S, Murakami M, Abe K, Maejima Y, Makino T.2010. Heavy metal contamination of agricultural soil and countermeasures in Japan.Paddy Water Environ, 8(3): 247-257. |
| [4] | Aziz R, Rafiq M T, Li T Q, Liu D, He Z L, Stoffella P J, Sun K W, Yang X E.2015. Uptake of cadmium by rice grown on contaminated soils and its bioavailability/toxicity in human cell lines (Caco-2/HL-7702).J Agric Food Chem, 63(13): 3599-3608. |
| [5] | Baysal C, Bortesi L, Zhu C, Farré G, Schillberg S, Christou P.2016. CRISPR/Cas9 activity in the rice OsBEIIb gene does not induce off-target effects in the closely related paralog OsBEIIa. Mol Breeding, 36: 108. |
| [6] | Clemens S, Aarts M G M, Thomine S, Verbruggen N.2013. Plant science: The key to preventing slow cadmium poisoning.Trends Plant Sci, 18(2): 92-99. |
| [7] | FAO/WHO.2018. Evaluation of certain food additives and contaminants (seventy-third report of the joint FAO/WHO expert committee on food additives). WHO Technical Report Series. |
| [8] | Grant C A, Clarke J M, Duguid S, Chaney R L.2008. Selection and breeding of plant cultivars to minimize cadmium accumulation.Sci Total Environ, 390: 301-310. |
| [9] | Hu L F, McBride M B, Cheng H, Wu J J, Shi J C, Xu J M, Wu L S.2011. Root-induced changes to cadmium speciation in the rhizosphere of two rice (Oryza sativa L.) genotypes. Environ Res, 111(3): 356-361. |
| [10] | Ishikawa S, Ishimaru Y, Igura M, Kuramata M, Abe T, Senoura T, Hase Y, Arao T, Nishizawa N K, Nakanishi H.2012. Ion-beam irradiation, gene identification, and marker-assisted breeding in the development of low-cadmium rice.Proc Natl Acad Sci USA, 109: 19166-19171. |
| [11] | Ishizaki T.2016. CRISPR/Cas9 in rice can induce new mutations in later generations, leading to chimerism and unpredicted segregation of the targeted mutation.Mol Breeding, 36: 165. |
| [12] | Jung C, Capistrano-Gossmann G, Braatz J, Sashidhar N, Melzer S.2017. Recent developments in genome editing and applications in plant breeding.Plant Breeding, 137(1): 1-9. |
| [13] | Lei Y, Lu L, Liu H Y, Li S, Xing F, Chen L L.2014. CRISPR-P: A web tool for synthetic single-guide RNA design of CRISPR- system in plants.Mol Plant, 7(9): 1494-1496. |
| [14] | Li S, Liu S M, Liu Y H, Lu H P, Tan Y Y, Huang J Z, Wei P C, Shu Q Y.2018. HRM-facilitated rapid identification and genotyping of mutations induced by CRISPR/Cas9 mutagenesis in rice.Crop Breeding Appl Biotechnol, 18: 184-191. |
| [15] | Li W L, Xu B B, Song Q J, Liu X M, Xu J M, Brookes P C.2014. The identification of ‘hotspots’ of heavy metal pollution in soil-rice systems at a regional scale in eastern China.Sci Total Environ, 472: 407-420. |
| [16] | Li W X, Wu S L, Liu Y H, Jin G L, Zhao H J, Fan L J, Shu Q Y.2016. Genome-wide profiling of genetic variation inAgrobacterium- transformed rice plants. J Zhejiang Univ Sci B, 17(12): 992-996. |
| [17] | Liu W Z, Xie X R, Ma X L, Li J, Chen J H, Liu Y G.2015. DSDecode: A web-based tool for decoding of sequencing chromatograms for genotyping of targeted mutations.Mol Plant, 8: 1431-1433. |
| [18] | Lu H P, Liu S M, Xu S L, Chen W Y, Zhou X, Tan Y Y, Huang J Z, Shu Q Y.2017. CRISPR-S: An active interference element for a rapid and inexpensive selection of genome-edited, transgene-free rice plants.Plant Biotechnol J, 15(11): 1371-1373. |
| [19] | Meharg A A, Norton G, Deacon C, Williams P, Adomako E E, Price A, Zhu Y G, Li G, Zhao F J, McGrath S, Villada A, Sommella A, de Silva P M C S, Brammer H, Dasgupta T, Islam M R.2013. Variation in rice cadmium related to human exposure.Environ Sci Technol, 47(11): 5613-5618. |
| [20] | Meng J, Zhong L B, Wang L, Liu X M, Tang C X, Chen H L, Xu J M.2018. Contrasting effects of alkaline amendments on the bioavailability and uptake of Cd in rice plants in a Cd- contaminated acid paddy soil.Environ Sci Pollut R, 25(9): 8827-8835. |
| [21] | Miyadate H, Adachi S, Hiraizumi A, Tezuka K, Nakazawa N, Kawamoto T, Katou K, Kodama I, Sakurai K, Takahashi H, Satoh-Nagasawa N, Watanabe A, Fujimura T, Akagi H.2010. OsHMA3, a P1B-type of ATPase affects root-to-shoot cadmium translocation in rice by mediating efflux into vacuoles.New Phytol, 189(1): 190-199. |
| [22] | Peng X X, Bai N N, Wang H H.2018. Isolation and expression profiles of cadmium stress-responsive rice WRKY15 transcription factor gene.Chin J Rice Sci, 32(2): 103-110. (in Chinese with English abstract) |
| [23] | Pinson S R M, Tarpley L, Yan W G, Yeater K, Lahner B, Yakubova E, Huang X Y, Zhang M, Guerinot M L, Salt D E.2015. Worldwide genetic diversity for mineral element concentrations in rice grain.Crop Sci, 55(1): 294-311. |
| [24] | Sasaki A, Yamaji N, Yokosho K, Ma J F.2012. Nramp5 is a major transporter responsible for manganese and cadmium uptake in rice.Plant Cell, 24(5): 2155-2167. |
| [25] | Sasaki A, Yamaji N, Ma J F.2014. Overexpression of OsHMA3 enhances Cd tolerance and expression of Zn transporter genes in rice. J Exp Bot, 65(20): 6013-6021. |
| [26] | Satoh-Nagasawa N, Mori M, Nakazawa N, Kawamoto T, Nagato Y, Sakurai K, Takahashi H, Watanabe A, Akagi H.2012. Mutations in rice (Oryza sativa) heavy metal ATPase 2(OsHMA2) restrict the translocation of zinc and cadmium. Plant Cell Physiol, 53(1): 213-224. |
| [27] | Shao G N, Xie L H, Jiao G A, Wei X J, Sheng Z H, Tang S Q, Hu P S.2017. CRISPR/CAS9-mediated editing of the fragrant gene Badh2 in rice. Chin J Rice Sci, 31: 216-222. (in Chinese with English abstract) |
| [28] | Shao J F, Fujii-Kashino M, Yamaji N, Fukuoka S, Shen R F, Ma J F.2017. Isolation and characterization of a rice line with high Cd accumulation for potential use in phytoremediation.Plant Soil, 410: 357-368. |
| [29] | Shimo H, Ishimaru Y, An G, Yamakawa T, Nakanishi H, Nishizawa N K.2011. Low cadmium (LCD), a novel gene related to cadmium tolerance and accumulation in rice.J Exp Bot, 62(15): 5727-5734. |
| [30] | Shu Q Y, Cui H R, Ye G Y, Wu D X, Xia Y W, Gao M W, Altosaar I.2002. Agronomic and morphological characterization of Agrobacterium-transformed Bt rice plants. Euphytica, 127(3): 345-352. |
| [31] | Smolders E, Mertens J.2013. Cadmium. In: Alloway B J. Heavy Metals in Soils: Trace Metals and Metalloids in Soils and Their Bioavailability. Dordrecht: Springer Netherlands: 283-311. |
| [32] | Sun Y L, Liu H M, Xu Q G.2017. Effects of cadmium stress on rice seed germination characteristics.Chin J Rice Sci, 31(4): 425-431. (in Chinese with English abstract) |
| [33] | Takahashi R, Ishimaru Y, Senoura T, Shimo H, Ishikawa S, Arao T, Nakanishi H, Nishizawa N K.2011. The OsNRAMP1 iron transporter is involved in Cd accumulation in rice.J Exp Bot, 62(14): 4843-4850. |
| [34] | Tang L, Mao B G, Li Y K, Lv Q M, Zhang L P, Chen C Y, He H J, Wang W P, Zeng X F, Shao Y, Pan Y L, Hu Y Y, Peng Y, Fu X Q, Li H Q, Xia S T, Zhao B R.2017. Knockout of OsNramp5 using the CRISPR/Cas9 system produces low Cd-accumulating indica rice without compromising yield. Sci Rep, 7(1): 14438. |
| [35] | Tang X J, Shen C F, Shi D Z, Cheema S A, Khan M I, Zhang C K, Chen Y X.2009. Heavy metal and persistent organic compound contamination in soil from Wenling: An emerging e-waste recycling city in Taizhou area, China.J Hazard Mater, 173: 653-600. |
| [36] | Ueno D, Yamaji N, Kono I, Huang C F, Ando T, Yano M, Ma J F.2010. Gene limiting cadmium accumulation in rice.Proc Natl Acad Sci USA, 107: 16500-16505. |
| [37] | Uraguchi S, Kamiya T, Sakamoto T, Kasai K, Sato Y, Nagamura Y, Yoshida A, Kyozuka J, Ishikawa S, Fujiwara T.2011. Low- affinity cation transporter (OsLCT1) regulates cadmium transport into rice grains.Proc Natl Acad Sci USA, 108: 20959-20964. |
| [38] | Uraguchi S, Kamiya T, Clemens S, Fujiwara T.2014. Characterization of OsLCT1, a cadmium transporter from indica rice(Oryza sativa). Physiol Plant, 151(3): 339-347. |
| [39] | Wright A V, Nuñez J K, Doudna J A.2016. Biology and applications of CRISPR systems: Harnessing nature’s toolbox for genome engineering.Cell, 164: 29-44. |
| [40] | Xu R F, Li H, Qin R Y, Wang L, Li L, Wei P C, Yang J B.2014. Gene targeting using the Agrobacterium tumefaciens-mediated CRISPR-Cas system in rice. Rice, 7(1): 5. |
| [41] | Yamaji N, Xia J X, Mitani-Ueno N, Yokosho K, Ma J F.2013. Preferential delivery of zinc to developing tissues in rice is mediated by P-type heavy metal ATPase OsHMA2.Plant Physiol, 162(2): 927-939. |
| [42] | Yao W Y, Sun L, Zhou H, Yang F, Mao D H, Wang J R, Chen L H, Zhang G Y, Dai J P, Xiao G Y, Chen C Y.2015. Additive, dominant parental effects control the inheritance of grain cadmium accumulation in hybrid rice.Mol Breeding, 35(1): 39. |
| [43] | Zeng F R, Mao Y, Cheng W D, Wu F B, Zhang G P.2008. Genotypic and environmental variation in chromium, cadmium and lead concentrations in rice.Environ Pollut, 153(2): 309-314. |
| [44] | Zhang H L, Huang J Z, Chen X Y, Tan Y Y, Shu Q Y.2014. Competitive amplification of differentially melting amplicons facilitates efficient genotyping of photoperiod- and temperature- sensitive genic male sterility in rice.Mol Breeding, 34(4): 1765-1776. |
| [45] | Zhang J S, Zhang H, Botella J R, Zhu J K.2017. Generation of new glutinous rice by CRISPR/Cas9-targeted mutagenesis of the Waxy gene in elite rice varieties. J Integr Plant Biol, 60(5): 369-375. |
| [46] | Zhang X Q, Zhang G P, Guo L B, Wang H Z, Zeng D L, Dong G J, Qian Q, Xue D W.2011. Identification of quantitative trait loci for Cd and Zn concentrations of brown rice grown in Cd-polluted soils.Euphytica, 180(2): 173-179. |
| [47] | Zhao F J, Ma Y B, Zhu Y G, Tang Z, McGrath S P.2015. Soil contamination in China: Current status and mitigation strategies.Environ Sci Technol, 49(2): 750-759. |
| [48] | Zhu H H, Chen C, Xu C, Zhu Q H, Huang D Y.2016. Effects of soil acidification and liming on the phytoavailability of cadmium in paddy soils of central subtropical China.Environ Pollut, 219: 99-106. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||