
Rice Science ›› 2020, Vol. 27 ›› Issue (5): 396-404.DOI: 10.1016/j.rsci.2020.03.004
• Orginal Article • Previous Articles Next Articles
Peng Qin1, Luchang Deng1,2, Weilan Chen1, Juan Huang3, Shijun Fan1, Bin Tu1, Jun Tan2, Hua Yuan1, Yuping Wang1, Bingtian Ma1, Shigui Li1( )
)
Received:2019-11-06
Accepted:2020-03-25
Online:2020-09-28
Published:2020-09-28
Contact:
Shigui Li
Peng Qin, Luchang Deng, Weilan Chen, Juan Huang, Shijun Fan, Bin Tu, Jun Tan, Hua Yuan, Yuping Wang, Bingtian Ma, Shigui Li. A Fragment Substitution in Promoter of MS92/PTC1 Causes Male Sterility in Rice[J]. Rice Science, 2020, 27(5): 396-404.
Add to citation manager EndNote|Ris|BibTeX
| Name | Primer sequence | Purpose | Gene symbol |
|---|---|---|---|
| RM24431 | CTTACTTAGCACGTGCCTGTTGC | Map-based cloning | |
| CATTGGGATCATGCATCTCTGC | |||
| RM3600 | TGCCCACACATGATGAGC | ||
| AACGGGCAAGAGATCTTCTG | |||
| RM7424 | CAGATCAAGCTAGCCACACAGC | ||
| GAAGGCAGAGCAGGAGAGAAGC | |||
| RM7048 | CGTGAAAGTGACGAGTTTCAGTCC | ||
| CGAAGTGAACATGGCACAAACC | |||
| Promoter-lost | GTGGGGGACACTCTTGGCTGACA | ||
| GGGTAGCCGGGCTGGCAGAA | |||
| MS92-GFP F | TCCCCCCGGGATGGCGCCTAAGATGGTGATCAG | Vector construction | |
| MS92-GFP R | GGACTAGTACAGTTGAAGGACGGGAACG | ||
| MS92-BD F | CGGAATTCATGGCGCCTAAGATGGTGATCAG | ||
| MS92-BD R | TTTGGATCCACAGTTGAAGGACGGGAACG | ||
| MS92ΔN1-BD F | CGGAATTCATGGCGCCTAAGATGGTGATCAG | ||
| MS92ΔN1-BD R | TTTGGATCCGCAGAGCGGCATGGAGCCGAG | ||
| MS92ΔN2-BD F | CACCGAATTCATGGCGCCTAAGATGGTGA | ||
| MS92ΔN2-BD R | TTTGGATCCGCGCTCGCCGCCGCCGCTCCCCT | ||
| MS92ΔN3-BD F | CACCGAATTCGTGCTGGTGCCGCACCTGTCGTGCT | ||
| MS92ΔN3-BD R | TTTGGATCCACAGTTGAAGGACGGGAACG | ||
| MS92ΔC1-BD F | CACCGAATTCGTCGTGGACTGCGCGTGCGG | ||
| MS92ΔC1-BD R | TTTGGATCCACAGTTGAAGGACGGGAACGACACGACG | ||
| MS92ΔC2-BD F | ACGCCATGCGCTTGCCGTCGTCGTC | ||
| MS92ΔC2-BD R | GACGACGACGGCAAGCGCATGGCGT | ||
| MS92ΔC3-BD F | GTGCTGCCACGCCTCCGCGATGTCGCAGCACGC | ||
| MS92ΔC3-BD R | GCGTGCTGCGACATCGCGGAGGCGTGGCAGCAC | ||
| MS92ΔC4-BD F | CCCGGCGCCCCGCGTGTTCTGCCACG | ||
| MS92ΔC4-BD R | CGTGGCAGAACACGCGGGGCGCCGGG | ||
| MS92 PHD GST F | TTTGGATCCGTCGTGGACTGCGCGTGCGG | ||
| MS92 PHD GST R | CCGGAATTCCGGACAGTTGAAGGACGGGAACG | ||
| RNA in situ F | CGTTGATTGGCAGCAACTAGC | ||
| RNA in situ R | CATATCAGATGCCTCCCCCAC | ||
| MS92 qPCR-F | GGCATCCCCACATCAACCT | qRT-PCR analysis | |
| MS92 qPCR-R | ATCTCGCTGAACACCCTCT | ||
| UBQ qPCR-F | GGAAGTAAGGAAGGAGGAG | ||
| UBQ qPCR-R | CAGAGGTGATGCTAAGGT | ||
| LOC_Os04g57490 qPCR-F | CAAGGAGTATCCCGTCTGCAA | OsCP1 | |
| LOC_Os04g57490 qPCR-R | AGAGATTGAATCAGGTTGTTCCG | ||
| LOC_Os07g05150 qPCR-F | AAGGAGGTTGCCGACGACAC | ||
| LOC_Os07g05150 qPCR-R | CGAAGCTCTTCAGCTCCTTCTC | ||
| LOC_Os06g02019 qPCR-F | CTGACCCCAAAATGTTCAACCC | OsKAO | |
| LOC_Os06g02019 qPCR-R | CTCACCCTGCACTTTGGATTTG | ||
| LOC_Os06g16300 qPCR-F | CTGATCGAGATCAAGAACCTGCT | ||
| LOC_Os06g16300 qPCR-R | AGAATAGTCTCAAGATTGGTCAGGT | ||
| LOC_Os04g51070 qPCR-F | TGAGGACGAACGAGATGTGATT | EAT1 | |
| LOC_Os04g51070 qPCR-R | TCACCTACTATTGAGGCCCTGT | ||
| LOC_Os09g27940 qPCR-F | CCACAGCTGCGTGAGGCCCGACACC | ||
| LOC_Os09g27940 qPCR-R | TCATCGTGCCGCAGCCGAACCCGAG | ||
| LOC_Os08g43290 qPCR-F | GAACAGGCTGCTGGCGTGCCGCGCGTA | OsC4 | |
| LOC_Os08g43290 qPCR-R | AGCAGTTGATCTGGCTGAGGTGGCAT | ||
| LOC_Os09g10260 qPCR-F | ATGTCGAAGCTAGTGAGCAACA | ||
| LOC_Os09g10260 qPCR-R | TTGTCGACATCGATGAAGCTGG | ||
| LOC_Os07g05180 qPCR-F | CGACACAATCCTCCTAATCAGC | ||
| LOC_Os07g05180 qPCR-R | GTTCTTGTCCACATCGATGAAC | ||
| LOC_Os07g23570 qPCR-F | CTCCACCTTCCTCACTGTTCAA | ||
| LOC_Os07g23570 qPCR-R | CAACCCAAGTAGGTCATTCCCA | ||
| LOC_Os10g20470 qPCR-F | TGTTCTTGCTTCGCAATGTCTG | ||
| LOC_Os10g20470 qPCR-R | ATGCCTACAAGGTAGAATGCCC | ||
| LOC_Os01g34970 qPCR-F | AACATTGCTGAACAGACTATTG | ||
| LOC_Os01g34970 qPCR-R | AACACCTTCTTGTAGAGTAGATT | ||
| LOC_Os07g42490 qPCR-F | GCTGGAAACAAGGAAACTGTGG | ||
| LOC_Os07g42490 qPCR-R | GAGGTGAGCCTCTTCTGTGATT |
Supplemental Table 1 Primers used in this study.
| Name | Primer sequence | Purpose | Gene symbol |
|---|---|---|---|
| RM24431 | CTTACTTAGCACGTGCCTGTTGC | Map-based cloning | |
| CATTGGGATCATGCATCTCTGC | |||
| RM3600 | TGCCCACACATGATGAGC | ||
| AACGGGCAAGAGATCTTCTG | |||
| RM7424 | CAGATCAAGCTAGCCACACAGC | ||
| GAAGGCAGAGCAGGAGAGAAGC | |||
| RM7048 | CGTGAAAGTGACGAGTTTCAGTCC | ||
| CGAAGTGAACATGGCACAAACC | |||
| Promoter-lost | GTGGGGGACACTCTTGGCTGACA | ||
| GGGTAGCCGGGCTGGCAGAA | |||
| MS92-GFP F | TCCCCCCGGGATGGCGCCTAAGATGGTGATCAG | Vector construction | |
| MS92-GFP R | GGACTAGTACAGTTGAAGGACGGGAACG | ||
| MS92-BD F | CGGAATTCATGGCGCCTAAGATGGTGATCAG | ||
| MS92-BD R | TTTGGATCCACAGTTGAAGGACGGGAACG | ||
| MS92ΔN1-BD F | CGGAATTCATGGCGCCTAAGATGGTGATCAG | ||
| MS92ΔN1-BD R | TTTGGATCCGCAGAGCGGCATGGAGCCGAG | ||
| MS92ΔN2-BD F | CACCGAATTCATGGCGCCTAAGATGGTGA | ||
| MS92ΔN2-BD R | TTTGGATCCGCGCTCGCCGCCGCCGCTCCCCT | ||
| MS92ΔN3-BD F | CACCGAATTCGTGCTGGTGCCGCACCTGTCGTGCT | ||
| MS92ΔN3-BD R | TTTGGATCCACAGTTGAAGGACGGGAACG | ||
| MS92ΔC1-BD F | CACCGAATTCGTCGTGGACTGCGCGTGCGG | ||
| MS92ΔC1-BD R | TTTGGATCCACAGTTGAAGGACGGGAACGACACGACG | ||
| MS92ΔC2-BD F | ACGCCATGCGCTTGCCGTCGTCGTC | ||
| MS92ΔC2-BD R | GACGACGACGGCAAGCGCATGGCGT | ||
| MS92ΔC3-BD F | GTGCTGCCACGCCTCCGCGATGTCGCAGCACGC | ||
| MS92ΔC3-BD R | GCGTGCTGCGACATCGCGGAGGCGTGGCAGCAC | ||
| MS92ΔC4-BD F | CCCGGCGCCCCGCGTGTTCTGCCACG | ||
| MS92ΔC4-BD R | CGTGGCAGAACACGCGGGGCGCCGGG | ||
| MS92 PHD GST F | TTTGGATCCGTCGTGGACTGCGCGTGCGG | ||
| MS92 PHD GST R | CCGGAATTCCGGACAGTTGAAGGACGGGAACG | ||
| RNA in situ F | CGTTGATTGGCAGCAACTAGC | ||
| RNA in situ R | CATATCAGATGCCTCCCCCAC | ||
| MS92 qPCR-F | GGCATCCCCACATCAACCT | qRT-PCR analysis | |
| MS92 qPCR-R | ATCTCGCTGAACACCCTCT | ||
| UBQ qPCR-F | GGAAGTAAGGAAGGAGGAG | ||
| UBQ qPCR-R | CAGAGGTGATGCTAAGGT | ||
| LOC_Os04g57490 qPCR-F | CAAGGAGTATCCCGTCTGCAA | OsCP1 | |
| LOC_Os04g57490 qPCR-R | AGAGATTGAATCAGGTTGTTCCG | ||
| LOC_Os07g05150 qPCR-F | AAGGAGGTTGCCGACGACAC | ||
| LOC_Os07g05150 qPCR-R | CGAAGCTCTTCAGCTCCTTCTC | ||
| LOC_Os06g02019 qPCR-F | CTGACCCCAAAATGTTCAACCC | OsKAO | |
| LOC_Os06g02019 qPCR-R | CTCACCCTGCACTTTGGATTTG | ||
| LOC_Os06g16300 qPCR-F | CTGATCGAGATCAAGAACCTGCT | ||
| LOC_Os06g16300 qPCR-R | AGAATAGTCTCAAGATTGGTCAGGT | ||
| LOC_Os04g51070 qPCR-F | TGAGGACGAACGAGATGTGATT | EAT1 | |
| LOC_Os04g51070 qPCR-R | TCACCTACTATTGAGGCCCTGT | ||
| LOC_Os09g27940 qPCR-F | CCACAGCTGCGTGAGGCCCGACACC | ||
| LOC_Os09g27940 qPCR-R | TCATCGTGCCGCAGCCGAACCCGAG | ||
| LOC_Os08g43290 qPCR-F | GAACAGGCTGCTGGCGTGCCGCGCGTA | OsC4 | |
| LOC_Os08g43290 qPCR-R | AGCAGTTGATCTGGCTGAGGTGGCAT | ||
| LOC_Os09g10260 qPCR-F | ATGTCGAAGCTAGTGAGCAACA | ||
| LOC_Os09g10260 qPCR-R | TTGTCGACATCGATGAAGCTGG | ||
| LOC_Os07g05180 qPCR-F | CGACACAATCCTCCTAATCAGC | ||
| LOC_Os07g05180 qPCR-R | GTTCTTGTCCACATCGATGAAC | ||
| LOC_Os07g23570 qPCR-F | CTCCACCTTCCTCACTGTTCAA | ||
| LOC_Os07g23570 qPCR-R | CAACCCAAGTAGGTCATTCCCA | ||
| LOC_Os10g20470 qPCR-F | TGTTCTTGCTTCGCAATGTCTG | ||
| LOC_Os10g20470 qPCR-R | ATGCCTACAAGGTAGAATGCCC | ||
| LOC_Os01g34970 qPCR-F | AACATTGCTGAACAGACTATTG | ||
| LOC_Os01g34970 qPCR-R | AACACCTTCTTGTAGAGTAGATT | ||
| LOC_Os07g42490 qPCR-F | GCTGGAAACAAGGAAACTGTGG | ||
| LOC_Os07g42490 qPCR-R | GAGGTGAGCCTCTTCTGTGATT |
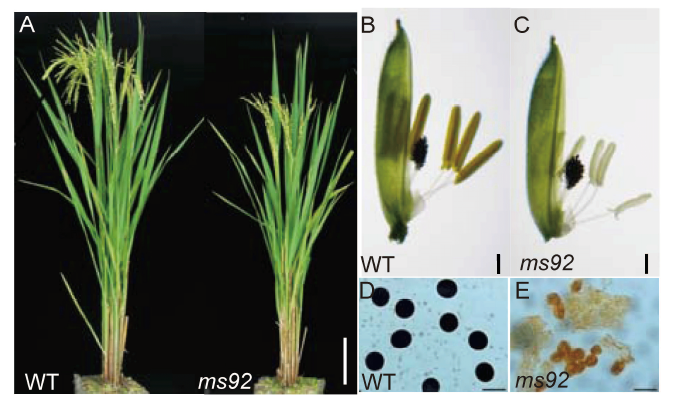

Fig. 1. Phenotypic comparisons of wild type (WT) and ms92 mutant.A, Comparison of WT and ms92 plants at the heading stage. Scale bar, 10 cm. B, WT floret at the mature pollen stage. Scale bar, 0.5 mm. C, ms92 floret at the mature pollen stage. Scale bar, 0.5 mm. D, Pollen of WT. Scale bar, 30 μm. E, Pollen of ms92. Scale bar, 30 μm.
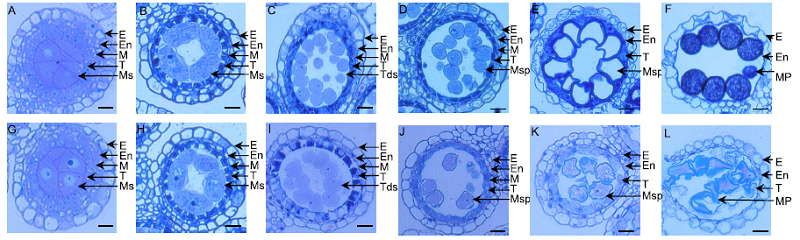
Fig. 2. Comparison of anther semi-thin section between wild type (WT) and ms92.A, WT at the microspore mother cell stage. B, WT at the meiosis stage. C, WT at the tetrad stage. D, WT at the young microspore stage. E, WT at the vacuolated pollen stage. F, WT at the mature pollen stage. G, ms92 at the microspore mother cell stage. H, ms92 at the meiosis stage. I, ms92 at the tetrad stage. J, ms92 at the young microspore stage. K, ms92 at the vacuolated pollen stage. L, ms92 at the mature pollen stage. Scale bars, 15 μm.E, Epidermis; En, Endothecium; M, Middle layer; MP, Mature pollen; Ms, Microsporocyte; Msp, Microspore; T, Tapetum; Tds, Tetrads.
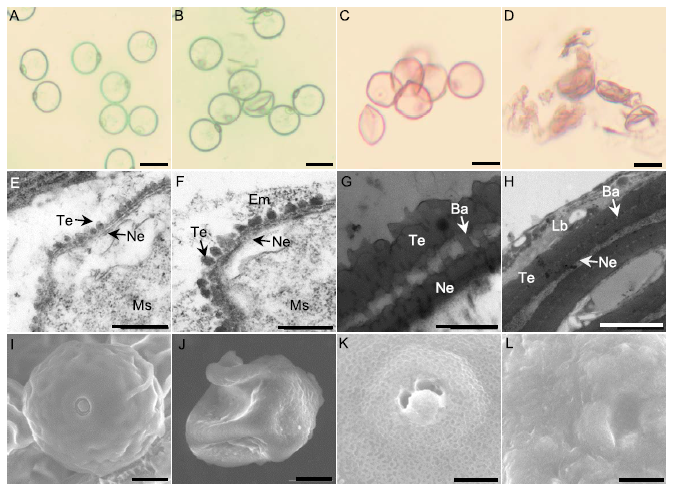
Fig. 3. Defective pollen wall of ms92.A and B, Microspore of wild type (A) and ms92 (B) at the young microspore stage treated with water. C and D, Microspore of wild type (C) and ms92 (D) at the young microspore stage treated with acetolysis mixture for 10 min. E and F, Primexine of wild type (E) and ms92 (F) at the young microspore stage. G and H, Exine of wild type (G) and ms92 (H) at the mature pollen stage. I and G, Pollen of wild type (I) and ms92 (J) at the mature pollen stage by scanning electron microscopy (SEM). K and L, Magnification of wild type (K) and ms92 (L) pollen.Scale bars are 20 μm in A to D, 0.5 μm in E to J, 2 μm in K and L.Ba, Baculum; Em, Electron-dense material; Lb, Lipid body; Ms, Microspore; Ne, Nexine; Te, Tectum.
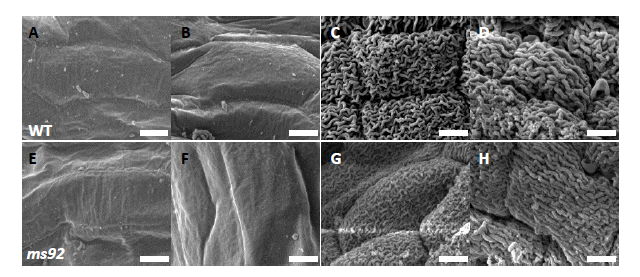
Supplemental Fig. 1. SEM analysis of anther surface of WT (A to D) and ms92 (E to H).AA and E, The meiosis stage. B and F, The young microspore stage. C and G, The vacuolated pollen stage; D and H, The mature pollen stage. Bars, 5 μm.
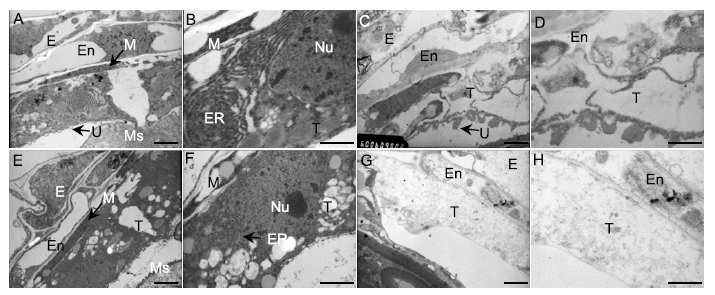
Fig. 4. Transmission electron microscopy analysis of wild type (WT) and ms92 anthers.A, Anthers of WT at the young microspore stage. Scale bar, 1 μm. B, Magnification of WT tapetum in A. Scale bar, 0.5 μm. C, Anthers of WT at the mature pollen stage. Scale bar, 1 μm. D, Magnification of WT tapetum in C. Scale bar, 0.5 μm. E, Anthers of ms92 at the young microspore stage. Scale bar, 1 μm. F, Magnification of ms92 tapetum in E. Scale bar, 0.5 μm. G, Anthers of ms92 at the mature pollen stage. Scale bar, 1 μm. H, Magnification of ms92 tapetum in G. Scale bar, 0.5 μm.E, Epidermis; En, Endothecium; M, Middle layer; Ms, Microsporocyte; Nu, Nucleus; ER, Endoplasmic reticulum; T, Tapetum; U, Ubisch bodies.
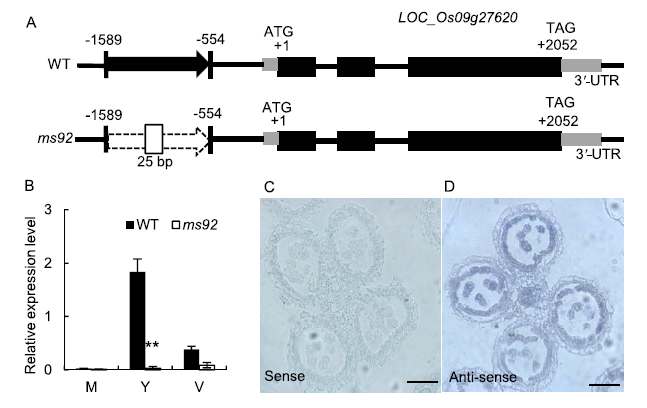
Fig. 5. Map based cloning of MS92, its expression pattern and RNA in situ hybridization.A, Schematic represents the exon (solid black box), intron (black lines) and UTR (gray lines) of MS92, 1 036 bp DNA fragment (black arrow) was replaced by 25 bp DNA fragment (white box) in the promoter. B, qRT-PCR analysis of MS92 expression in wild type (WT) and ms92 spikelet. M, Meiosis stage; Y, Young microspore stage; V, Vacuolated pollen stage. Values are Mean ± SD (n = 3). C, RNA in situ hybridization analysis of MS92 using WT anther sections at the young microspore stage with sense probe. Scale bar, 50 μm. D, RNA in situ hybridization analysis of MS92 using WT anther sections at the young microspore stage with anti-sense probe. Scale bar, 50 μm.
Fertile | sterile | χ2(1:1) | |
|---|---|---|---|
ptc1×MS92 B | 17 | 12 | 0.209 |
ms92×PTC1 B | 22 | 26 | 0.606 |
Supplemental Table 2 Allelic analysis between ms92 and ptc1.
Fertile | sterile | χ2(1:1) | |
|---|---|---|---|
ptc1×MS92 B | 17 | 12 | 0.209 |
ms92×PTC1 B | 22 | 26 | 0.606 |
| Gene ID | Gene symble | Expression tissue a | Function | LOGms92/WT | Isolation | Reference |
|---|---|---|---|---|---|---|
| LOC_Os04g48400 | HTH1 | Epidermis-expressed | Anther cuticle and Pollen development | -3.07 | KD and M | Xu et al, 2017 |
| LOC_Os03g18480 | OsTDF1 | Tapetum-specific | Tapetum degeneration | 1.15 | M | Cai et al, 2015 |
| LOC_Os04g37570 | OsAP37 | Tapetum-specific | Tapetum degeneration | -2.76 | None | Niu et al, 2013 |
| LOC_Os04g51070 | EAT1/DTD1 | Tapetum-specific | Tapetum degeneration | -1.93 | M | Niu et al, 2013 |
| LOC_Os08g43290 | OsC4 | Tapetum-specific | Pollen exine development | -1.81 | None | Tsuchiya et al, 1994 |
| LOC_Os01g02190 | OsNIP4;2 | RPMCT | Pollen development and pollination | -4.99 | None | Pérez et al, 2016 |
| LOC_Os10g38050 | OsNP1 | RPMCT | Anther cuticle and Pollen exine patterning | -2.64 | M | Liu et al, 2017 |
| LOC_Os12g13930 | KAR | RPMCT | Pollen exine formation | -3.65 | None | Aya et al, 2009 |
| LOC_Os01g62430 | OsERG1 | RPMCT | Plant defense | -2.23 | None | Kim et al, 2003 |
| LOC_Os04g45960 | OsSub42 | RPMCT | Panicle development | -3.03 | None | Sharma et al, 2012 |
| LOC_Os03g07140 | OsMS2/DPW | Tapetum and microspore | Pollen wall development | -2.56 | M | Shi et al, 2011 |
| LOC_Os04g57490 | OsCP1 | Tapetum and microspore | Pollen development | -1.87 | KD | Lee et al, 2004 |
| LOC_Os05g49900 | OsCHS | Tapetum and microspore | Pollen exine formation(decreased) | -3.13 | None | Zhang et al, 2008 |
| LOC_Os06g47880 | OsCDKG1 | Tapetum and microspore | Pollen wall formation | -2.78 | None | Huang et al, 2013 |
| LOC_Os04g52320 | OsQRT3 | Tapetum and microspore | Pollen mother cell wall degradation | -2.11 | None | Rhee et al, 2003 |
| LOC_Os04g02640 | OsKCS6 | Tapetum and microspore | Pollen wall development | -1.33 | None | A et al, 2013 |
| LOC_Os01g50622 | OsBBD1 | Tapetum and microspore | Callose deposition in leaf | -1.29 | None | You et al, 2010 |
| LOC_Os12g04980 | DMC1A/DMC1 | Tapetum and microspore | Homologous paring in rice meiosis | 1.82 | KD and M | Deng and Wang, 2007 |
| LOC_Os01g50940 | OsMYL1 | Tapetum and microspore | Anther cell identity | 1.51 | None | Yang et al, 2016 |
| LOC_Os08g06110 | OsLHY | Tapetum and microspore | Flowering time regulation | 1.11 | None | Ogiso et al, 2010 |
| LOC_Os01g15900 | RDD1 | Tapetum and microspore | Phytochrome regulation | 1.09 | KD | Iwamoto et al, 2009 |
| LOC_Os05g50930 | OsSIG5 | Tapetum and microspore | Light-responsive gene | 1.17 | None | Kubota et al, 2007 |
| LOC_Os07g48830 | WSI76 | Tapetum and microspore | Stress-induced gene | 1.45 | None | Rabbani et al, 2003 |
| LOC_Os04g51150 | OsAOX1a | Tapetum and microspore | Cold tolerance | 1.25 | OX | Li et al, 2013 |
| LOC_Os11g32650 | CHS | Tapetum and microspore | Plant defense | -2.38 | None | Qiu et al, 2007 |
| LOC_Os05g51090 | OsSWEET5 | Tapetum and microspore | Crosstalk between sugar and auxin | -6.79 | OX | Zhou et al, 2014 |
| LOC_Os01g64890 | OsMGT1 | Tapetum and microspore | Salt or aluminum tolerance | -1.52 | M | Chen et al, 2017 |
| LOC_Os07g37090 | OSRIP18 | Tapetum and microspore | Salt and drought tolerance | -3.46 | OX | Jiang et al, 2012 |
| LOC_Os05g35400 | OsBiP4 | Tapetum and microspore | Endoplasmic reticulum stress response | -1.81 | KD | Wakasa et al, 2012 |
| LOC_Os04g48350 | OsDREB1-1 | Tapetum and microspore | Stress response | -1.29 | None | Tian et al, 2005 |
| LOC_Os04g46940 | OsHMA5 | Tapetum and microspore | Xylem loading of copper | -1.35 | M | Deng et al, 2013 |
Supplemental Table 3 Summary of 31 known genes with changed expression in both ms92 and ptc1.
| Gene ID | Gene symble | Expression tissue a | Function | LOGms92/WT | Isolation | Reference |
|---|---|---|---|---|---|---|
| LOC_Os04g48400 | HTH1 | Epidermis-expressed | Anther cuticle and Pollen development | -3.07 | KD and M | Xu et al, 2017 |
| LOC_Os03g18480 | OsTDF1 | Tapetum-specific | Tapetum degeneration | 1.15 | M | Cai et al, 2015 |
| LOC_Os04g37570 | OsAP37 | Tapetum-specific | Tapetum degeneration | -2.76 | None | Niu et al, 2013 |
| LOC_Os04g51070 | EAT1/DTD1 | Tapetum-specific | Tapetum degeneration | -1.93 | M | Niu et al, 2013 |
| LOC_Os08g43290 | OsC4 | Tapetum-specific | Pollen exine development | -1.81 | None | Tsuchiya et al, 1994 |
| LOC_Os01g02190 | OsNIP4;2 | RPMCT | Pollen development and pollination | -4.99 | None | Pérez et al, 2016 |
| LOC_Os10g38050 | OsNP1 | RPMCT | Anther cuticle and Pollen exine patterning | -2.64 | M | Liu et al, 2017 |
| LOC_Os12g13930 | KAR | RPMCT | Pollen exine formation | -3.65 | None | Aya et al, 2009 |
| LOC_Os01g62430 | OsERG1 | RPMCT | Plant defense | -2.23 | None | Kim et al, 2003 |
| LOC_Os04g45960 | OsSub42 | RPMCT | Panicle development | -3.03 | None | Sharma et al, 2012 |
| LOC_Os03g07140 | OsMS2/DPW | Tapetum and microspore | Pollen wall development | -2.56 | M | Shi et al, 2011 |
| LOC_Os04g57490 | OsCP1 | Tapetum and microspore | Pollen development | -1.87 | KD | Lee et al, 2004 |
| LOC_Os05g49900 | OsCHS | Tapetum and microspore | Pollen exine formation(decreased) | -3.13 | None | Zhang et al, 2008 |
| LOC_Os06g47880 | OsCDKG1 | Tapetum and microspore | Pollen wall formation | -2.78 | None | Huang et al, 2013 |
| LOC_Os04g52320 | OsQRT3 | Tapetum and microspore | Pollen mother cell wall degradation | -2.11 | None | Rhee et al, 2003 |
| LOC_Os04g02640 | OsKCS6 | Tapetum and microspore | Pollen wall development | -1.33 | None | A et al, 2013 |
| LOC_Os01g50622 | OsBBD1 | Tapetum and microspore | Callose deposition in leaf | -1.29 | None | You et al, 2010 |
| LOC_Os12g04980 | DMC1A/DMC1 | Tapetum and microspore | Homologous paring in rice meiosis | 1.82 | KD and M | Deng and Wang, 2007 |
| LOC_Os01g50940 | OsMYL1 | Tapetum and microspore | Anther cell identity | 1.51 | None | Yang et al, 2016 |
| LOC_Os08g06110 | OsLHY | Tapetum and microspore | Flowering time regulation | 1.11 | None | Ogiso et al, 2010 |
| LOC_Os01g15900 | RDD1 | Tapetum and microspore | Phytochrome regulation | 1.09 | KD | Iwamoto et al, 2009 |
| LOC_Os05g50930 | OsSIG5 | Tapetum and microspore | Light-responsive gene | 1.17 | None | Kubota et al, 2007 |
| LOC_Os07g48830 | WSI76 | Tapetum and microspore | Stress-induced gene | 1.45 | None | Rabbani et al, 2003 |
| LOC_Os04g51150 | OsAOX1a | Tapetum and microspore | Cold tolerance | 1.25 | OX | Li et al, 2013 |
| LOC_Os11g32650 | CHS | Tapetum and microspore | Plant defense | -2.38 | None | Qiu et al, 2007 |
| LOC_Os05g51090 | OsSWEET5 | Tapetum and microspore | Crosstalk between sugar and auxin | -6.79 | OX | Zhou et al, 2014 |
| LOC_Os01g64890 | OsMGT1 | Tapetum and microspore | Salt or aluminum tolerance | -1.52 | M | Chen et al, 2017 |
| LOC_Os07g37090 | OSRIP18 | Tapetum and microspore | Salt and drought tolerance | -3.46 | OX | Jiang et al, 2012 |
| LOC_Os05g35400 | OsBiP4 | Tapetum and microspore | Endoplasmic reticulum stress response | -1.81 | KD | Wakasa et al, 2012 |
| LOC_Os04g48350 | OsDREB1-1 | Tapetum and microspore | Stress response | -1.29 | None | Tian et al, 2005 |
| LOC_Os04g46940 | OsHMA5 | Tapetum and microspore | Xylem loading of copper | -1.35 | M | Deng et al, 2013 |
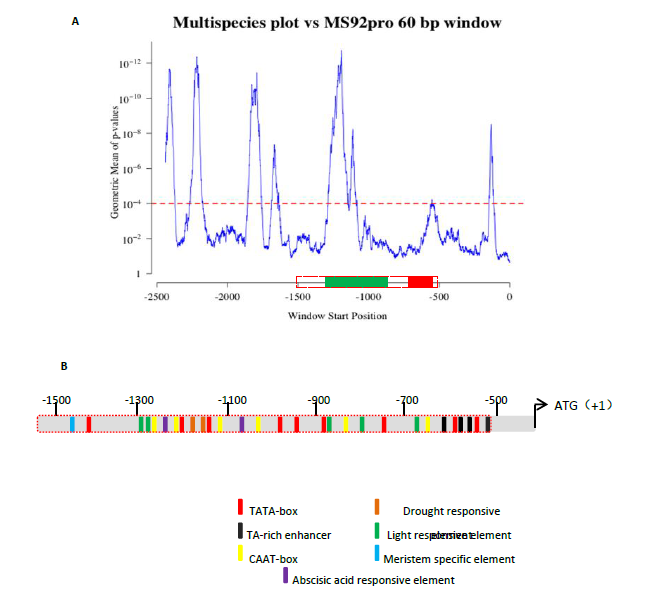
Fig. 3. Conservation plots and cis-elements analyses of the substitution sequence in the promoter of MS92. A, Comparison of conservation plots relative to individual plant species. The promoter of the MS92 was compared with that of orthologous genes from 30 plant species using a 60-bp window and a step width of 1 bp. The substitution sequence was denoted with a red dotted box, including conserved core region (red box) and responsive region (green box).B, Graphical representation of 1.5-kb upstream regulatory region of MS92 and presence of key cis-elements. Square box represents the presence of important cis-regulatory motif on the sequence with respect to its TSS.
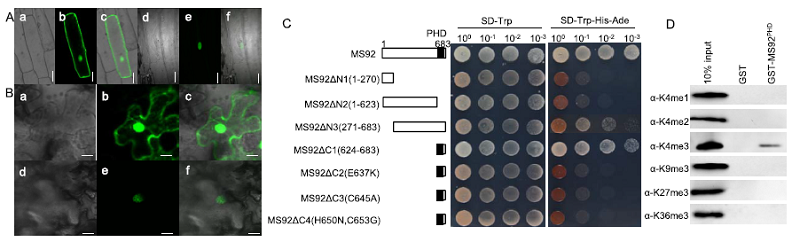
Fig. 6. Subcellular localization and transcriptional activation assay of MS92 protein.A, Subcellular localization of MS92 protein in onion epidermis cells. (a) Bright-filed image of 35S::GFP; (b) Green fluorescence signal of 35S::GFP; (c) Merged image of 35S::GFP; (d) Bright-filed image of MS92::GFP; (e) Green fluorescence signal of MS92::GFP; (f) Merged image of MS92::GFP. Scale bars, 20 μm. B, Subcellular localization of MS92 protein in Nicotiana benthamiana leaf cells. (a) Bright-filed image of 35S::GFP; (b) Green fluorescence signal of 35S::GFP; (c) Merged image of 35S::GFP; (d) Bright-filed image of MS92::GFP; (e) Green fluorescence signal of MS92::GFP; (f) Merged image of MS92::GFP. Scale bars, 20 μm. C, Transcriptional activation assay of MS92 protein and truncated proteins. PHD, Plant homeodomain. D, Pull-down assay of calf histones with MS92 PHD and analyzed by western blots using the antibodies indicated on the left. GST, Only glutathione-S-transferases tagged protein. GST-MS92PHD, PHD tagged with GST protein.
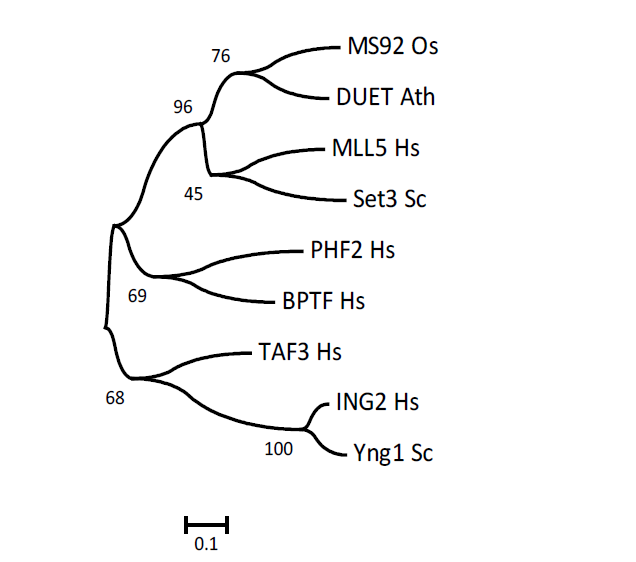
Supplemental Fig. 4. Neighbor-joining tree using PHD (Plant Homeodomain) finger protein sequences.Hs, Homo sapiens; Sc, Saccharomyces cerevisiae, Ath, Arabidopsis thaliana; Os, Oryza sativa.The number for each interior branch shows the percentage of the bootstrap value (1000 replicates). The scale bar indicates the estimated number of amino acid substitutions per site.
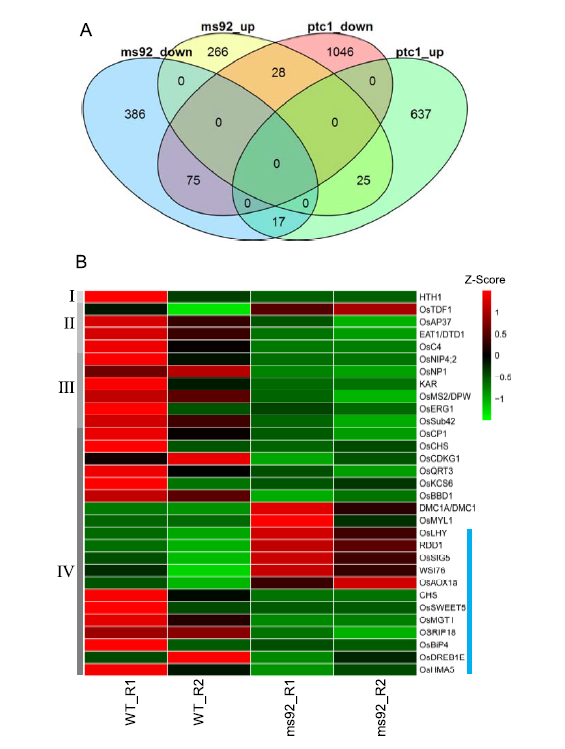
Fig. 7. Differential expression gene analysis between using RNA-seq of wild type (WT) and ms92 and microarray data of WT and ptc1.A, Venn diagram depicting the numbers of DEG between WT and ms92 or ptc1. B, Cluster analysis of 31 genes altered in both ms92 and ptc1 which had been reported. Genes were classified into Clusters I, II, III and IV based on their expression patterns. Other details are showed in Supplemental Table 2. Red and green represents higher and lower expression levels, respectively. Two independent biological samples were analyzed in heatmap.
| [1] | Ariizumi T, Toriyama K.2011. Genetic regulation of sporopollenin synthesis and pollen exine development. Annu Rev Plant Biol, 62(1): 437-460. |
| [2] | Aya K, Ueguchi-Tanaka M, Kondo M, Hamada K, Yano K, Nishimura M, Matsuoka M.2009. Gibberellin modulates anther development in rice via the transcriptional regulation of GAMYB. Plant Cell, 21(5): 1453-1472. |
| [3] | Cai C F, Zhu J, Lou Y, Guo Z L, Xiong S X, Wang K, Yang Z N.2015. The functional analysis of OsTDF1 reveals a conserved genetic pathway for tapetal development between rice and Arabidopsis. Sci Bull, 60(12): 1073-1082. |
| [4] | Chen J B, He F, Qin P, Wang Y P, Xu J, Li Y Y, Zheng J, Hu J T, Li S G.2010. Genetic analysis and gene mapping of a rice recessive male sterile mutant. Plant Breeding, 129(3): 313-317. |
| [5] | Deng L C, Zhang S W, Wang G L, Fan S J, Li M, Chen W L, Tu B, Tan J, Wang Y P, Ma B T, Li S G, Qin P.2017. Down-regulation of OsEMF2b caused semi-sterility due to anther and pollen development defects in rice. Front Plant Sci, 8: 1998. |
| [6] | Dityatkin S Y, Lisovskaya K V, Panzhava N N, Iliashenko B N.1972. Frozen-thawed bacteria as recipients of isolated coliphage DNA. Biochim Biophys Acta: Nucl Acids Protein Synth, 281(3): 319-323. |
| [7] | Du X, Fei Y Y, Wang F Q, Xu Y, Wang J, Li W Q, Zhao L,Chen Z H, Liang G H, Zhou Y, Yang J.2019. Thermo-sensitive male sterile line created by editing TMS5 gene in japonica rice. Chin J Rice Sci, 33(5): 429-435. (in Chinese with English abstract) |
| [8] | Fernández Gomez J, Wilson Z A.2014. A barley PHD finger transcription factor that confers male sterility by affecting tapetal development. Plant Biotechnol J, 12(6): 765-777. |
| [9] | Hobo T, Suwabe K, Aya K, Suzuki G, Yano K, Ishimizu T, Fujita M, Kikuchi S, Hamada K, Miyano M, Fujioka T, Kaneko F, Kazama T, Mizuta Y, Takahashi H, Shiono K, Nakazono M, Tsusumi N, Naamura Y, Kurata N, Watanabe M, Matsuoka M.2008. Various spatiotemporal expression profiles of anther-expressed genes in rice. Plant Cell Physiol, 49(10): 1417-1428. |
| [10] | Huang M, Zhang L.1999. Association of the movement protein of alfalfa mosaic virus with the endoplasmic reticulum and its trafficking in epidermal cells of onion bulb scales. Mol Plant Microbe Interact, 12(8): 680-690. |
| [11] | Ito T, Nagata N, Yoshiba Y, Ohme-Takagi M, Ma H, Shinozaki K.2007. Arabidopsis MALE STERILITY1 encodes a PHD-type transcription factor and regulates pollen and tapetum development. Plant Cell, 19(11): 3549-3562. |
| [12] | Lee S, Jung K H, An G, Chung Y Y.2004. Isolation and characterization of a rice cysteine protease gene, OsCP1, using T-DNA gene-trap system. Plant Mol Biol, 54(5): 755-765. |
| [13] | Lee W Y, Lee D, Chung W, Kwon C S.2009. Arabidopsis ING and Alfin1-like protein families localize to the nucleus and bind to H3K4me3/2 via plant homeodomain fingers. Plant J, 58(3): 511-524. |
| [14] | Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, de Peer Y V, Rouzé P, Rombauts S.2002. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucl Acids Res, 30(1): 325-327. |
| [15] | Li H, Yuan Z, Vizcay-Barrena G, Yang C, Liang W Q, Zong J, Wilson Z A, Zhang D B.2011. PERSISTENT TAPETAL CELL1 encodes a PHD-finger protein that is required for tapetal cell death and pollen development in rice. Plant Physiol, 156(2): 615-630. |
| [16] | Lisch D.2013. How important are transposons for plant evolution? Nat Rev Genet, 14(1): 49-61. |
| [17] | Louveaux J, Maurizio A, Vorwohl G.1978. Methods of melissopalynology. Bee World, 59(4): 139-157. |
| [18] | Ma X Z, Sun X Q, Li C M, Huan R, Sun C H, Wang Y, Xiao F L, Wang Q, Chen P R, Ma F R, Zhang K, Wang P R, Deng X J.2017. Map-based cloning and characterization of the novel yellow-green leaf gene ys83 in rice (Oryza sativa). Plant Physiol Biochem, 111: 1-9. |
| [19] | Malovannaya A, Lanz R B, Jung S Y, Bulynko Y, Le N T, Chan D W, Ding C, Shi Y, Yucer N, Krenciute G, Kim B J, Li C S, Chen R, Li W, Wang Y, O’Malley B W, Qin J.2011. Analysis of the human endogenous coregulator complexome. Cell, 145(5): 787-799. |
| [20] | Mouriz A, Lopez-Gonzalez L, Jarillo J A, Pineiro M.2015. PHDs govern plant development. Plant Signal Behav, 10(7): e993253. |
| [21] | Niu N N, Liang W Q, Yang X J, Jin W L, Wilson Z A, Hu J P, Zhang D B.2013. EAT1 promotes tapetal cell death by regulating aspartic proteases during male reproductive development in rice. Nat Commun, 4: 1445. |
| [22] | Picot E, Krusche P, Tiskin A, Carré I, Ott S.2010. Evolutionary analysis of regulatory sequences (EARS) in plants. Plant J, 64(1): 165-176. |
| [23] | Qin P, Tu B, Wang Y P, Deng L C, Quilichini T D, Li T, Wang H, Ma B T, Li S G.2013. ABCG15 encodes an ABC transporter protein, and is essential for post-meiotic anther and pollen exine development in rice. Plant Cell Physiol, 54(1): 138-154. |
| [24] | Robinson M D, McCarthy D J, Smyth G K.2010. edgeR: A bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics, 26(1): 139-140. |
| [25] | Sorensen A M, Kröber S, Unte U S, Huijser P, Dekker K, Saedler H.2003. The Arabidopsis ABORTED MICROSPORES (AMS) gene encodes a MYC class transcription factor. Plant J, 33(2): 413-423. |
| [26] | Wilson Z A, Morroll S M, Dawson J, Swarup R, Tighe P J.2001. The Arabidopsis MALE STERILITY1 (MS1) gene is a transcriptional regulator of male gametogenesis, with homology to the PHD- finger family of transcription factors. Plant J, 28(1): 27-39. |
| [27] | Yang C Y, Vizcay-Barrena G, Conner K, Wilson Zoe A.2007. MALE STERILITY1 is required for tapetal development and pollen wall biosynthesis. Plant Cell, 19(11): 3530-3548. |
| [28] | Yang Z F, Liu L, Sun L P, Yu P, Zhang P P, Abbas A, Xiang X J, Wu W X, Zhang Y X, Cao L Y, Cheng S H.2019. OsMS1 functions as a transcriptional activator to regulate programmed tapetum development and pollen exine formation in rice. Plant Mol Biol, 99: 175-191. |
| [29] | Zhang D S, Liang W Q, Yin C, Zong J, Gu F, Zhang D B.2010. OsC6, encoding a lipid transfer protein, is required for postmeiotic anther development in rice. Plant Physiol, 154(1): 149-162. |
| [30] | Zhang W, Sun Y J, Timofejeva L, Chen C B, Grossniklaus U, Ma H.2006. Regulation of Arabidopsis tapetum development and function by DYSFUNCTIONAL TAPETUM1 (DYT1) encoding a putative bHLH transcription factor. Development, 133(16): 3085-3095. |
| [31] | Zhang Z B, Zhu J, Gao J F, Wang C, Li H, Li H, Zhang H Q, Zhang S, Wang D M, Wang Q X.2007. Transcription factor AtMYB103 is required for anther development by regulating tapetum development, callose dissolution and exine formation in Arabidopsis. Plant J, 52(3): 528-538. |
| [32] | Zhu J, Chen H, Li H, Gao J F, Jiang H, Wang C, Guan Y F, Yang Z N.2008. Defective in tapetal development and function 1 is essential for anther development and tapetal function for microspore maturation in Arabidopsis. Plant J, 55(2): 266-277. |
| [33] | Zhu J, Lou Y, Xu X F, Yang Z N.2011. A genetic pathway for tapetum development and function in Arabidopsis. J Integr Plant Biol, 53(11): 892-900. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [13] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||