
Rice Science ›› 2020, Vol. 27 ›› Issue (5): 414-422.DOI: 10.1016/j.rsci.2020.04.008
• Orginal Article • Previous Articles Next Articles
Hui Wang, Jiayu Zhang, Farkhanda Naz, Juan Li, Shuangfei Sun, Guanghua He, Ting Zhang, Yinghua Ling, Fangming Zhao( )
)
Received:2019-12-05
Accepted:2020-04-15
Online:2020-09-28
Published:2020-09-28
Hui Wang, Jiayu Zhang, Farkhanda Naz, Juan Li, Shuangfei Sun, Guanghua He, Ting Zhang, Yinghua Ling, Fangming Zhao. Identification of Rice QTLs for Important Agronomic Traits with Long-Kernel CSSL-Z741 and Three SSSLs[J]. Rice Science, 2020, 27(5): 414-422.
Add to citation manager EndNote|Ris|BibTeX
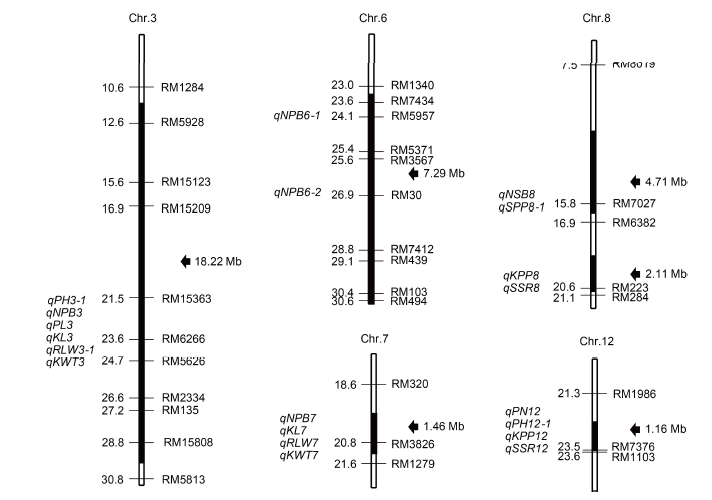
Fig. 1. Chromosome substitution segments of Z741 and QTLs identified in them.Physical distances (Mb) and mapped QTLs are marked to the left of each chromosome. Markers and lengths of the substitution segments are displayed to the right. Black sections on each chromosome indicate the substitution segments.KL, Kernel length; KPP, Number of kernel per panicle; KWT, 1000-kernel weight; NPB, Number of primary branches per plant; NSB, Number of secondary branches per plant; PH, Plant height; PL, Panicle length; PN, Number of panicles per plant; RLW, Ratio of length to width; SPP, Number of spikelets per panicle, SSR, Seed-setting rate.

Supplemental Fig. 1. Phenotypes of Nipponbare and Z741.A, Whole-plant phenotype of Nipponbare and Z741 at the milky stage. B, Kernel shape of Nipponbare and Z741.
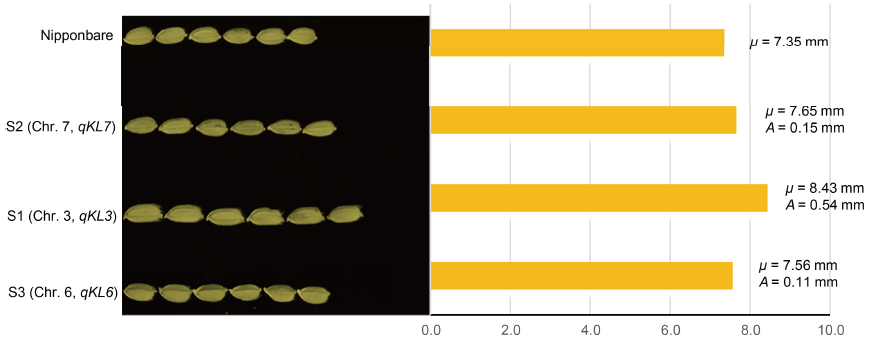
Fig. 2. Kernel lengths of Nipponbare and three single-segment substitution lines S1, S2 and S3.Left, Images of aligned kernels for lines of given genotypes; Right, Statistics of kernel length. μ, Mean value of kernel lengths; A, Additive effect value for each QTL controlling kernel length.
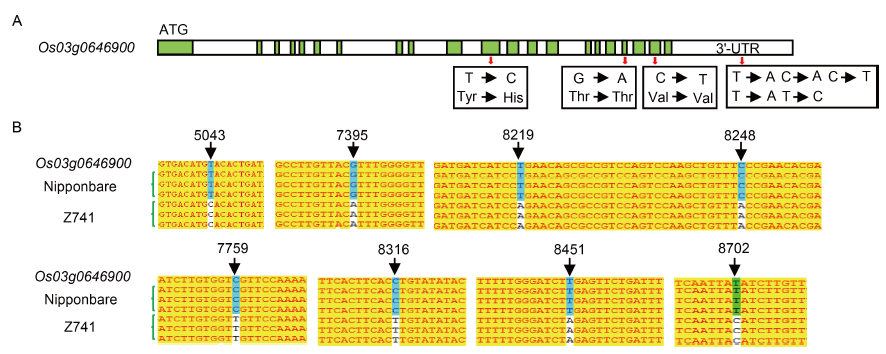
Fig. 3. Comparison of OsPPKL1 sequences in Nipponbare and Z741.A, Sketch map of OsPPKL1 (Os03g0646900) gene. Green boxes refer to exons, white boxes refer to introns and 3′-UTR. Red arrows indicate the alteration of a codon. B, Different sequences of OsPPKL1 between Nipponbare and Z741. Yellow indicates conserved sequence, and black arrows mark mutation sites.
| [1] | Bai X F, Luo L J, Yan W H, Kovi M R, Zhan W, Xing Y Z.2010. Genetic dissection of rice grain shape using a recombinant inbred line population derived from two contrasting parents and fine mapping a pleiotropic quantitative trait locus qGL7. BMC Genet, 11: 16. |
| [2] | Balakrishnan D, Surapaneni M, Mesapogu S, Neelamraju S.2019. Development and use of chromosome segment substitution lines as a genetic resource for crop improvement. Theor Appl Genet, 132: 1-25. |
| [3] | Ding Z Q, Lin Z F, Li Q, Wu H, Xiang C Y, Wang J F.2015. DNL1 encodes cellulose synthase-like D4 is a major QTL for plant height and leaf width in rice (Oryza sativa L.). Biochem Biophys Res Commun, 457(2): 133-140. |
| [4] | Eshed Y, Zamir D.1995. An introgression line population of Lycopersicon pennellii in the cultivated tomato enables the identification and fine mapping of yield-associated QTL. Genetics, 141(3): 1147-1162. |
| [5] | Furuta T, Uehara K, Angeles-Shim R B, Shim J, Ashikari M, Takashi T.2014. Development and evaluation of chromosome segment substitution lines (CSSLs) carrying chromosome segments derived from Oryza rufipogon in the genetic background of Oryza sativa L. Breeding Sci, 63(5): 468-475. |
| [6] | Gao X Y, Zhang J Q, Zhang X J, Zhou J, Jiang Z S, Huang P, Tang Z B, Bao Y M, Cheng J P, Tang H J, Zhang W H, Zhang H S, Huang J.2019. Rice qGL3/OsPPKL1 functions with the GSK3/ SHAGGY-like kinase OsGSK3 to modulate brassinosteroid signaling. Plant Cell, 31(5): 1077-1093. |
| [7] | Han Z M, Hu W, Tan C, Xing Y Z.2017. QTLs for heading date and plant height under multiple environments in rice. Genetica, 145: 67-77. |
| [8] | Hong Z, Ueguchi-Tanaka M, Shimizu-Sato S, Inukai Y, Fujioka S, Shimada Y, Takatsuto S, Agetsuma M, Yoshida S, Watanabe Y, Uozu S, Kitano H, Ashikari M, Matsuoka M.2002. Loss-of- function of a rice brassinosteroid biosynthetic enzyme, C-6 oxidase, prevents the organized arrangement and polar elongation of cells in the leaves and stem. Plant J, 32(4): 495-508. |
| [9] | Hu Z J, He H H, Zhang S Y, Sun F, Xin X Y, Wang W X, Qian X, Yang J S, Luo X J.2012. A kelch motif-containing serine/ threonine protein phosphatase determines the large grain QTL trait in rice. J Integr Plant Biol, 54(12): 979-990. |
| [10] | Ishimaru K, Hirotsu N, Madoka Y, Murakami N, Hara N, Onodera H, Kashiwagi T, Ujiie K, Shimizu B I, Onishi A, Miyagawa H, Katoh E.2013. Loss of function of the IAA-glucose hydrolase gene TGW6 enhances rice grain weight and increases yield. Nat Genet, 45(6): 707-711. |
| [11] | Li M M, Xu L, Ren J F, Cao G L, Yu L Q, He H H, Han L Z, Koh H J.2010. Identification of quantitative trait loci for grain traits in japonica rice. Agric Sci China, 9(7): 929-936. |
| [12] | Li Z H, Riaz A, Zhang Y X, Anis G B, Zhu A K, Cao L Y, Cheng S H.2019. Quantitative trait loci mapping for rice yield-related traits using chromosomal segment substitution lines. Rice Sci, 26(5): 261-264. |
| [13] | Liu G F, Zhang Z M, Zhu H T, Zhao F M, Ding X H, Zeng R Z, Li W T, Zhang G Q.2008. Detection of QTLs with additive effects and additive-by-environment interaction effects on panicle number in rice (Oryza sativa L.) with single-segment substitution lines. Theor Appl Genet, 116: 923-931. |
| [14] | Liu Q, Han R X, Wu K, Zhang J Q, Ye Y F, Wang S S, Chen J F, Pan Y J, Li Q, Xu X P, Zhou J W, Tao D Y, Wu Y J, Fu X D.2018. G-protein βγ subunits determine grain size through interaction with MADS-domain transcription factors in rice. Nat Commun, 9: 852. |
| [15] | Ma F Y, Zhu X Y, Wang H, Wang S M, Cui G Q, Zhang T, Yang Z L, He G H, Ling Y H, Wang N, Zhao F M.2019. Identification of QTL for kernel number-related traits in a rice chromosome segment substitution line and fine mapping of qSP1. Crop J, 7(4): 494-503. |
| [16] | McCouch S R, Kochert G, Yu Z H, Wang Z Y, Khush G S, Coffman W R, Tanksley S D.1988. Molecular mapping of rice chromosomes. Theor Appl Genet, 76(6): 815-829. |
| [17] | Paterson A H, Damon S, Hewitt J D, Zamir D, Rabinowitch H D, Lincoln S E, Lander E S, Tanksley S D.1991. Mendelian factors underlying quantitative traits in tomato-comparison across species generations and environments. Genetics, 127(1): 181-197. |
| [18] | Qi P, Lin Y S, Song X J, Shen J B, Huang W, Shan J X, Zhu M Z, Jiang L W, Gao J P, Lin H X.2012. The novel quantitative trait locus GL3.1 controls rice grain size and yield by regulating cyclin-T1;3. Cell Res, 22(12): 1666-1680. |
| [19] | Shao G N, Tang S Q, Luo J, Jiao G A, Wei X G, Tang A, Wu J L, Zhuang J Y, Hu P S.2010. Mapping of qGL7-2, a grain length QTL on chromosome 7 of rice. J Genet Genom, 37(8): 523-531. |
| [20] | Song X J, Huang W, Shi M, Zhu M Z, Lin H X.2007. A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat Genet, 39: 623-630. |
| [21] | Song X J, Kuroha T, Ayano M, Furuta T, Nagai K, Komeda N, Segami S, Miura K, Ogawa D, Kamura T, Suzuki T, Higashiyama T, Yamasaki M, Mori H, Inukai Y, Wu J Z, Kitano H, Sakakibara H, Jacobsen S E, Ashikari M.2015. Rare allele of a previously unidentified histone H4 acetyltransferase enhances grain weight, yield, and plant biomass in rice. Proc Natl Acad Sci USA, 112(1): 76-81. |
| [22] | Wang J, Zhu J Y, Zhou Y, Yang J, Wang Z D, Fan F J, Liang G H, Zhong W G.2012. Mapping of QTLs for panicle length using CSSLs in rice (Oryza sativa L.). Acta Agric Sin, 27(1): 68-73. (in Chinese with English abstract) |
| [23] | Wang S K, Wu K, Yuan Q B, Liu X Y, Liu Z B, Lin X Y, Zeng R Z, Zhu H T, Dong G J, Qian Q, Zhang G Q, Fu X D.2012. Control of grain size, shape and quality by OsSPL16 in rice. Nat Genet, 44(8): 950-954. |
| [24] | Wang S K, Li S, Liu Q, Wu K, Zhang J Q, Wang S S, Wang Y, Chen X B, Zhang Y, Gao C X, Wang F, Huang H X, Fu X D.2015. The OsSPL16-GW7 regulatory module determines grain shape and simultaneously improves rice yield and grain quality. Nat Genet, 47(8): 949-954. |
| [25] | Wang Y X, Xiong G S, Hu J, Jiang L, Yu H, Xu J, Fang Y X, Zeng L J, Xu E B, Xu J, Ye W J, Meng X B, Liu R F, Chen H Q, Jing Y H, Wang Y H, Zhu X D, Li J Y, Qian Q.2015. Copy number variation at the GL7 locus contributes to grain size diversity in rice. Nat Genet, 47(8): 944-948. |
| [26] | Xiang J, Li Y, Fan Y W, Xu J H, Zheng L Y, He G H, Yang Z L, Wang N, Zhao F M.2015. Identification and morphological analysis of a rice chromosome segment substitution line carrying a major effect gene for late heading date and mapping of Ehd4-2. Acta Agron Sin, 41(5): 683-691. (in Chinese with English abstract) |
| [27] | Yang W F, Zhan P L, Lin S J, Gou Y J, Zhang G Q, Wang S K.2019. Research progress of grain shape genetics in rice. J South China Agric Univ, 40(5): 203-210. (in Chinese with English abstract) |
| [28] | Zhang X J, Wang J F, Huang J, Lan H X, Wang C L, Yin C F, Wu Y Y, Tang H J, Qian Q, Li J Y, Zhang H S.2012. Rare allele of OsPPKL1 associated with grain length causes extra-large grain and a significant yield increase in rice. Proc Natl Acad Sci USA, 109: 21534-21539. |
| [29] | Zhang Y S, Luo L J, Xu C G, Zhang Q F, Xing Y Z.2006. Quantitative trait loci for panicle size, heading date and plant height co-segregating in trait-performance derived near-isogenic lines of rice (Oryza sativa). Theor Appl Genet, 113(2): 361-368. |
| [30] | Zhao D S, Li Q F, Zhang C Q, Zhang C, Yang Q Q, Pan L X, Ren X Y, Lu J, Gu M H, Liu Q Q.2018. GS9 acts as a transcriptional activator to regulate rice grain shape and appearance quality. Nat Commun, 9: 1240. |
| [31] | Zhao F M, Zhu H T, Ding X H, Zeng R Z, Zhang Z M, Li W T, Zhang G Q.2007. Detection of QTLs for important agronomic traits and analysis of their stabilities using SSSLs in rice. Agric Sci China, 6(7): 769-778. |
| [32] | Zhao F M, Zhang G Q, Yang Z L, He G H.2014. Pyramiding QTL for yield-related traits and grain shape in rice using single- segment substitution lines. Ind J Genet Plant Breeding, 74(4): 496-498. |
| [33] | Zhao F M, Zhu H T, Zeng R Z, Zhang G Q, Xu S Z.2016. Detection of additive and additive × environment interaction effects of QTLs for yield component traits of rice using single segment substitution lines (SSSL). Plant Breeding, 135(4): 452-458. |
| [1] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [2] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [3] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [4] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [5] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [6] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [7] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [8] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [9] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [10] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [14] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| [15] | Lu Xuedan, Li Fan, Xiao Yunhua, Wang Feng, Zhang Guilian, Deng Huabing, Tang Wenbang. Grain Shape Genes: Shaping the Future of Rice Breeding [J]. Rice Science, 2023, 30(5): 379-404. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||