
Rice Science ›› 2020, Vol. 27 ›› Issue (2): 113-123.DOI: 10.1016/j.rsci.2020.01.002
• Research Paper • Previous Articles Next Articles
B. ANGELES-SHIM Rosalyn1,2,4( ), P. REYES Vincent1, M. del VALLE Marilyn1, S. LAPIS Ruby1, SHIM Junghyun1,4, SUNOHARA Hidehiko3, K. JENA Kshirod1, ASHIKARI Motoyuki2, DOI Kazuyuki3
), P. REYES Vincent1, M. del VALLE Marilyn1, S. LAPIS Ruby1, SHIM Junghyun1,4, SUNOHARA Hidehiko3, K. JENA Kshirod1, ASHIKARI Motoyuki2, DOI Kazuyuki3
Received:2018-09-18
Accepted:2019-01-15
Online:2020-03-28
Published:2019-11-28
B. ANGELES-SHIM Rosalyn, P. REYES Vincent, M. del VALLE Marilyn, S. LAPIS Ruby, SHIM Junghyun, SUNOHARA Hidehiko, K. JENA Kshirod, ASHIKARI Motoyuki, DOI Kazuyuki. Marker-Assisted Introgression of Quantitative Resistance Gene pi21 Confers Broad Spectrum Resistance to Rice Blast[J]. Rice Science, 2020, 27(2): 113-123.
Add to citation manager EndNote|Ris|BibTeX
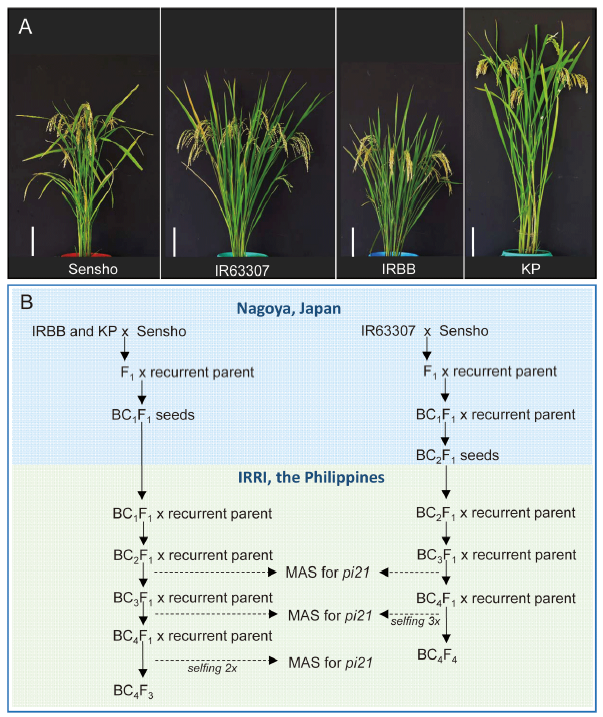
Fig. 1. Introgression of pi21 allele from Sensho to different rice genetic backgrounds. A, Gross morphology of Sensho and the recipient parents IR63307, IRBB and KP. Bar = 20 cm. B, Marker-assisted breeding scheme used to transfer the pi21 allele from Sensho to the different rice backgrounds. The blue panel indicates the initial crosses and backcrosses that were carried out in Nagoya, Japan, whereas the green panel indicates the marker-assisted backcrossing and generation advance that were conducted at International Rice Research Institute (IRRI) in the Philippines. MAS, Marker-assisted sclection.
| Isolate name | Groupb | Subgroupc | Philippine collection site |
|---|---|---|---|
| Ca89 | I | b | Luzon and Mindanao islands |
| IK81-3 | II | - | Southern Luzon island |
| M39-1-3-8-1 | III | a | Southern Luzon island |
| V850256 | III | c | Southern Luzon island |
| M64-1-3-9-1 | III | a | Southern Luzon island |
| V86010 | IV | e | Central and Southern Luzon island |
| BZ64-1 | IV | - | unknown |
| BN111 | V | a | Southern Luzon island |
| PO6-6 | V | b | Southern Luzon island |
| M39-1-2-21-2 | VI | b | Central and Southern Luzon island |
| JMB8401 | XII | a | Southern Luzon island |
| 43 | XVIII | - | unknown |
| M36-1-3-10-1 | XIX | b | Southern Luzon and Mindanao islands |
| 9475-1-3 | XX | - | unknown |
| BN209 | XXII | a | Southern Luzon island |
| M101-1-2-9-1 | XXII | a | Southern Luzon island |
| IK81-25 | XXIII | - | unknown |
| JMB840610 | XXIV | b | Southern Luzon island |
| 5167-1 | XIX | - | unknown |
| Pi9-G7-2K-1 | - | - | - |
Supplemental Table 1. Twenty blast isolates from the Philippines used in leaf blast isolate-specific screening of WISH lines carrying the pi21 allele.a
| Isolate name | Groupb | Subgroupc | Philippine collection site |
|---|---|---|---|
| Ca89 | I | b | Luzon and Mindanao islands |
| IK81-3 | II | - | Southern Luzon island |
| M39-1-3-8-1 | III | a | Southern Luzon island |
| V850256 | III | c | Southern Luzon island |
| M64-1-3-9-1 | III | a | Southern Luzon island |
| V86010 | IV | e | Central and Southern Luzon island |
| BZ64-1 | IV | - | unknown |
| BN111 | V | a | Southern Luzon island |
| PO6-6 | V | b | Southern Luzon island |
| M39-1-2-21-2 | VI | b | Central and Southern Luzon island |
| JMB8401 | XII | a | Southern Luzon island |
| 43 | XVIII | - | unknown |
| M36-1-3-10-1 | XIX | b | Southern Luzon and Mindanao islands |
| 9475-1-3 | XX | - | unknown |
| BN209 | XXII | a | Southern Luzon island |
| M101-1-2-9-1 | XXII | a | Southern Luzon island |
| IK81-25 | XXIII | - | unknown |
| JMB840610 | XXIV | b | Southern Luzon island |
| 5167-1 | XIX | - | unknown |
| Pi9-G7-2K-1 | - | - | - |
| Cultivar and line | Initial reading | Final reading | |||||
|---|---|---|---|---|---|---|---|
| SES | DLA (%) | Phenotype | SES | DLA (%) | Phenotype | ||
| Lijiangxintuanheigu (LTH, susceptible control) | 5.3 | 34 | MR | ND | ND | S | |
| CO39 (susceptible control) | 7.1 | 60 | S | 8.2 | 80 | S | |
| IR65482-4-136-2-2 (resistant control) | 1.4 | 1 | R | 1.6 | 1 | R | |
| Sensho | 1.1 | 2 | R | 1.5 | 2 | R | |
| IR63307-4B0-B-2 (IR63307) | 0.5 | 1 | R | 0.5 | 1 | R | |
| WISH110:1-1-11-12-1 | 1.0 | 1 | R | 1.0 | 1 | R | |
| WISH110:1-1-11-5-4 | 2.0 | 1 | R | 0.5 | 1 | R | |
| WISH110:2-5-2-1-1 | 1.0 | 1 | R | 0.5 | 1 | R | |
| WISH110:2-5-2-12-1 | 0.0 | 0 | R | 0.5 | 1 | R | |
| WISH110:2-5-2-22-1 | 0.5 | 1 | R | 0.5 | 1 | R | |
| IRBB4/5/13/21 (IRBB) | 4.0 | 5 | MR | 5.0 | 15 | MR | |
| WISH48:1-3-1-1 | 2.0 | 1 | R | 2.0 | 1 | R | |
| WISH48:1-3-18-1 | 2.0 | 3 | R | 3.5 | 5 | R | |
| WISH48:1-3-2-1 | 1.0 | 1 | R | 1.0 | 1 | R | |
| WISH48:1-3-20-1 | 3.0 | 3 | R | 1.0 | 1 | R | |
| WISH48:1-3-21-1 | 1.0 | 1 | R | 2.0 | 1 | R | |
| Kinandang Patong (KP) | 0.0 | 0 | R | 0.5 | 0 | R | |
| WISH40:1-3-16-11 | 0.0 | 0 | R | 0.5 | 0 | R | |
| WISH40:1-3-3-9 | 0.0 | 0 | R | 0.0 | 0 | R | |
| WISH40:1-3-7-7 | 0.0 | 0 | R | 0.0 | 0 | R | |
Table 1 Reaction of WISH lines to natural leaf blast infection in the field based on SES scores (IRRI, 2014).
| Cultivar and line | Initial reading | Final reading | |||||
|---|---|---|---|---|---|---|---|
| SES | DLA (%) | Phenotype | SES | DLA (%) | Phenotype | ||
| Lijiangxintuanheigu (LTH, susceptible control) | 5.3 | 34 | MR | ND | ND | S | |
| CO39 (susceptible control) | 7.1 | 60 | S | 8.2 | 80 | S | |
| IR65482-4-136-2-2 (resistant control) | 1.4 | 1 | R | 1.6 | 1 | R | |
| Sensho | 1.1 | 2 | R | 1.5 | 2 | R | |
| IR63307-4B0-B-2 (IR63307) | 0.5 | 1 | R | 0.5 | 1 | R | |
| WISH110:1-1-11-12-1 | 1.0 | 1 | R | 1.0 | 1 | R | |
| WISH110:1-1-11-5-4 | 2.0 | 1 | R | 0.5 | 1 | R | |
| WISH110:2-5-2-1-1 | 1.0 | 1 | R | 0.5 | 1 | R | |
| WISH110:2-5-2-12-1 | 0.0 | 0 | R | 0.5 | 1 | R | |
| WISH110:2-5-2-22-1 | 0.5 | 1 | R | 0.5 | 1 | R | |
| IRBB4/5/13/21 (IRBB) | 4.0 | 5 | MR | 5.0 | 15 | MR | |
| WISH48:1-3-1-1 | 2.0 | 1 | R | 2.0 | 1 | R | |
| WISH48:1-3-18-1 | 2.0 | 3 | R | 3.5 | 5 | R | |
| WISH48:1-3-2-1 | 1.0 | 1 | R | 1.0 | 1 | R | |
| WISH48:1-3-20-1 | 3.0 | 3 | R | 1.0 | 1 | R | |
| WISH48:1-3-21-1 | 1.0 | 1 | R | 2.0 | 1 | R | |
| Kinandang Patong (KP) | 0.0 | 0 | R | 0.5 | 0 | R | |
| WISH40:1-3-16-11 | 0.0 | 0 | R | 0.5 | 0 | R | |
| WISH40:1-3-3-9 | 0.0 | 0 | R | 0.0 | 0 | R | |
| WISH40:1-3-7-7 | 0.0 | 0 | R | 0.0 | 0 | R | |
| Line name | Blast isolates | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1a | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | |
| LTH | 3b/S | 3/S | 0/R | 3/S | 4/S | 4/S | 3/S | 4/S | 3/S | 3/S | 5/S | 5/S | 3/S | 5/S | 5/S | 5/S | 5/S | 5/S | 4/S | 4/S |
| CO39 | 4/S | 3/S | 2/R | 4/S | 4/S | 4/S | 3/S | 4/S | 4/S | 3/S | 5/S | 5/S | 4/S | 4/S | 4/S | 5/S | 5/S | 5/S | 4/S | 0/R |
| IR65482-4-136-2-2 | 0/R | 0/R | 0/R | 0/R | 0/R | 0/R | 0/R | 0/R | 0/R | 0/R | 1/R | 0/R | 4/S | 0/R | 0/R | 1/R | 2/R | 0/R | 0/R | 0/R |
| IR72 | 0/R | 0/R | 0/R | 1/R | 0/R | 3/S | 0/R | 0/R | 0/R | 0/R | 2/R | 1/R | 0/R | 0/R | 0/R | 0/R | 2/R | 1/R | 0/R | 2/R |
| Sensho | 0/R | 1/R | 0/R | 4/S | 2/R | 1/R | 1/R | 1/R | 3/S | 3/S | 2/R | 2/R | 1/R | 0/R | 2/R | 4/S | 2/R | 1/R | 1/R | 0/R |
| IR63307 | 2/R | 1/R | 0/R | 0/R | 0/R | 2/R | 1/R | 2/R | 2/R | 1/R | 1/R | 2/R | 2/R | 0/R | 5/S | 5/S | 2/R | 0/R | 4/S | 0/R |
| WISH 110:1-1-11-12-1 | 1/R | 0/R | 0/R | 0/R | 0/R | 1/R | 1/R | 0/R | 1/R | 0/R | 1/R | 1/R | 1/R | 0/R | 2/R | 2/R | 1/R | 0/R | 2/R | 0/R |
| WISH 110:1-1-11-5-4 | 1/R | 1/R | 0/R | 0/R | 0/R | 1/R | 1/R | 1/R | 1/R | 1/R | 1/R | 1/R | 1/R | 0/R | 2/R | 2/R | 1/R | 0/R | 1/R | 0/R |
| WISH 110:2-5-2-1-1 | 1/R | 0/R | 0/R | 0/R | 0/R | 1/R | 1/R | 1/R | 1/R | 1/R | 1/R | 1/R | 1/R | 0/R | 2/R | 2/R | 1/R | 0/R | 2/R | 0/R |
| WISH 110:2-5-2-12-1 | 1/R | 0/R | 0/R | 1/R | 0/R | 1/R | 0/R | 1/R | 1/R | 0/R | 1/R | 1/R | 1/R | 0/R | 1/R | 1/R | 1/R | 0/R | 2/R | 0/R |
| WISH 110:2-5-2-22-1 | 1/R | 1/R | 0/R | 0/R | 0/R | 1/R | 0/R | 1/R | 1/R | 0/R | 1/R | 1/R | 1/R | 0/R | 2/R | 2/R | 1/R | 0/R | 2/R | 0/R |
| IRBB | 2/R | 3/S | 1/R | 4/S | 0/R | 2/R | 3/S | 2/R | 0/R | 0/R | 5/S | 0/R | 4/S | 4/S | 2/R | 4/S | 5/S | 5/S | 0/R | 0/R |
| WISH 48:1-3-1-1 | 1/R | 1/R | 0/R | 3/S | 0/R | 1/R | 1/R | 0/R | 0/R | 0/R | 2/R | 0/R | 1/R | 2/R | 1/R | 4/S | 4/S | 2/R | 0/R | 0/R |
| WISH 48:1-3-18-1 | 1/R | 1/R | 0/R | 3/S | 0/R | 2/R | 2/R | 1/R | 0/R | 0/R | 2/R | 0/R | 1/R | 2/R | 1/R | 4/S | 4/S | 3/S | 1/R | 0/R |
| WISH 48:1-3-2-1 | 1/R | 1/R | 1/R | 1/R | 0/R | 0/R | 1/R | 1/R | 0/R | 0/R | 2/R | 0/R | 1/R | 1/R | 0/R | 3/S | 3/S | 2/R | 0/R | 0/R |
| WISH 48:1-3-20-1 | 1/R | 2/R | 1/R | 4/S | 0/R | 2/R | 1/R | 1/R | 0/R | 0/R | 3/S | 0/R | 1/R | 1/R | 1/R | 4/S | 4/S | 3/S | 0/R | 0/R |
| WISH 48:1-3-21-1 | 1/R | 1/R | 0/R | 3/S | ND | 0/R | 0/R | 1/R | 0/R | 0/R | 1/R | 0/R | 2/R | 1/R | 1/R | 4/S | 4/S | 2/R | 0/R | 0/R |
| KP | 1/R | 2/R | 0/R | 3/S | 4/S | 1/R | 1/R | 2/R | 4/S | 3/S | 3/S | 5/S | 1/R | 2/R | 4/S | 2/R | 3/S | 1/R | 4/S | 0/R |
| WISH 40:1-3-16-11 | 1/R | 0/R | 0/R | 2/R | 2/R | 1/R | 0/R | 1/R | 1/R | 0/R | 2/R | 2/R | 0/R | 1/R | 2/R | 1/R | 2/R | 1/R | 3/S | 1/R |
| WISH 40:1-3-3-9 | 1/R | 1/R | 0/R | 2/R | 2/R | 1/R | 0/R | 1/R | 1/R | 2/R | 1/R | 2/R | 0/R | 1/R | 2/R | 1/R | 1/R | 1/R | 1/R | 0/R |
| WISH 40:1-3-7-7 | 1/R | 0/R | 0/R | 1/R | 0/R | 1/R | 1/R | 0/R | 2/R | 1/R | 1/R | 2/R | 0/R | 1/R | 2/R | 1/R | 1/R | 1/R | 1/R | 0/R |
Supplemental Table 2. Reaction of WISH lines to twenty leaf blast isolates in the glasshouse.
| Line name | Blast isolates | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1a | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | |
| LTH | 3b/S | 3/S | 0/R | 3/S | 4/S | 4/S | 3/S | 4/S | 3/S | 3/S | 5/S | 5/S | 3/S | 5/S | 5/S | 5/S | 5/S | 5/S | 4/S | 4/S |
| CO39 | 4/S | 3/S | 2/R | 4/S | 4/S | 4/S | 3/S | 4/S | 4/S | 3/S | 5/S | 5/S | 4/S | 4/S | 4/S | 5/S | 5/S | 5/S | 4/S | 0/R |
| IR65482-4-136-2-2 | 0/R | 0/R | 0/R | 0/R | 0/R | 0/R | 0/R | 0/R | 0/R | 0/R | 1/R | 0/R | 4/S | 0/R | 0/R | 1/R | 2/R | 0/R | 0/R | 0/R |
| IR72 | 0/R | 0/R | 0/R | 1/R | 0/R | 3/S | 0/R | 0/R | 0/R | 0/R | 2/R | 1/R | 0/R | 0/R | 0/R | 0/R | 2/R | 1/R | 0/R | 2/R |
| Sensho | 0/R | 1/R | 0/R | 4/S | 2/R | 1/R | 1/R | 1/R | 3/S | 3/S | 2/R | 2/R | 1/R | 0/R | 2/R | 4/S | 2/R | 1/R | 1/R | 0/R |
| IR63307 | 2/R | 1/R | 0/R | 0/R | 0/R | 2/R | 1/R | 2/R | 2/R | 1/R | 1/R | 2/R | 2/R | 0/R | 5/S | 5/S | 2/R | 0/R | 4/S | 0/R |
| WISH 110:1-1-11-12-1 | 1/R | 0/R | 0/R | 0/R | 0/R | 1/R | 1/R | 0/R | 1/R | 0/R | 1/R | 1/R | 1/R | 0/R | 2/R | 2/R | 1/R | 0/R | 2/R | 0/R |
| WISH 110:1-1-11-5-4 | 1/R | 1/R | 0/R | 0/R | 0/R | 1/R | 1/R | 1/R | 1/R | 1/R | 1/R | 1/R | 1/R | 0/R | 2/R | 2/R | 1/R | 0/R | 1/R | 0/R |
| WISH 110:2-5-2-1-1 | 1/R | 0/R | 0/R | 0/R | 0/R | 1/R | 1/R | 1/R | 1/R | 1/R | 1/R | 1/R | 1/R | 0/R | 2/R | 2/R | 1/R | 0/R | 2/R | 0/R |
| WISH 110:2-5-2-12-1 | 1/R | 0/R | 0/R | 1/R | 0/R | 1/R | 0/R | 1/R | 1/R | 0/R | 1/R | 1/R | 1/R | 0/R | 1/R | 1/R | 1/R | 0/R | 2/R | 0/R |
| WISH 110:2-5-2-22-1 | 1/R | 1/R | 0/R | 0/R | 0/R | 1/R | 0/R | 1/R | 1/R | 0/R | 1/R | 1/R | 1/R | 0/R | 2/R | 2/R | 1/R | 0/R | 2/R | 0/R |
| IRBB | 2/R | 3/S | 1/R | 4/S | 0/R | 2/R | 3/S | 2/R | 0/R | 0/R | 5/S | 0/R | 4/S | 4/S | 2/R | 4/S | 5/S | 5/S | 0/R | 0/R |
| WISH 48:1-3-1-1 | 1/R | 1/R | 0/R | 3/S | 0/R | 1/R | 1/R | 0/R | 0/R | 0/R | 2/R | 0/R | 1/R | 2/R | 1/R | 4/S | 4/S | 2/R | 0/R | 0/R |
| WISH 48:1-3-18-1 | 1/R | 1/R | 0/R | 3/S | 0/R | 2/R | 2/R | 1/R | 0/R | 0/R | 2/R | 0/R | 1/R | 2/R | 1/R | 4/S | 4/S | 3/S | 1/R | 0/R |
| WISH 48:1-3-2-1 | 1/R | 1/R | 1/R | 1/R | 0/R | 0/R | 1/R | 1/R | 0/R | 0/R | 2/R | 0/R | 1/R | 1/R | 0/R | 3/S | 3/S | 2/R | 0/R | 0/R |
| WISH 48:1-3-20-1 | 1/R | 2/R | 1/R | 4/S | 0/R | 2/R | 1/R | 1/R | 0/R | 0/R | 3/S | 0/R | 1/R | 1/R | 1/R | 4/S | 4/S | 3/S | 0/R | 0/R |
| WISH 48:1-3-21-1 | 1/R | 1/R | 0/R | 3/S | ND | 0/R | 0/R | 1/R | 0/R | 0/R | 1/R | 0/R | 2/R | 1/R | 1/R | 4/S | 4/S | 2/R | 0/R | 0/R |
| KP | 1/R | 2/R | 0/R | 3/S | 4/S | 1/R | 1/R | 2/R | 4/S | 3/S | 3/S | 5/S | 1/R | 2/R | 4/S | 2/R | 3/S | 1/R | 4/S | 0/R |
| WISH 40:1-3-16-11 | 1/R | 0/R | 0/R | 2/R | 2/R | 1/R | 0/R | 1/R | 1/R | 0/R | 2/R | 2/R | 0/R | 1/R | 2/R | 1/R | 2/R | 1/R | 3/S | 1/R |
| WISH 40:1-3-3-9 | 1/R | 1/R | 0/R | 2/R | 2/R | 1/R | 0/R | 1/R | 1/R | 2/R | 1/R | 2/R | 0/R | 1/R | 2/R | 1/R | 1/R | 1/R | 1/R | 0/R |
| WISH 40:1-3-7-7 | 1/R | 0/R | 0/R | 1/R | 0/R | 1/R | 1/R | 0/R | 2/R | 1/R | 1/R | 2/R | 0/R | 1/R | 2/R | 1/R | 1/R | 1/R | 1/R | 0/R |
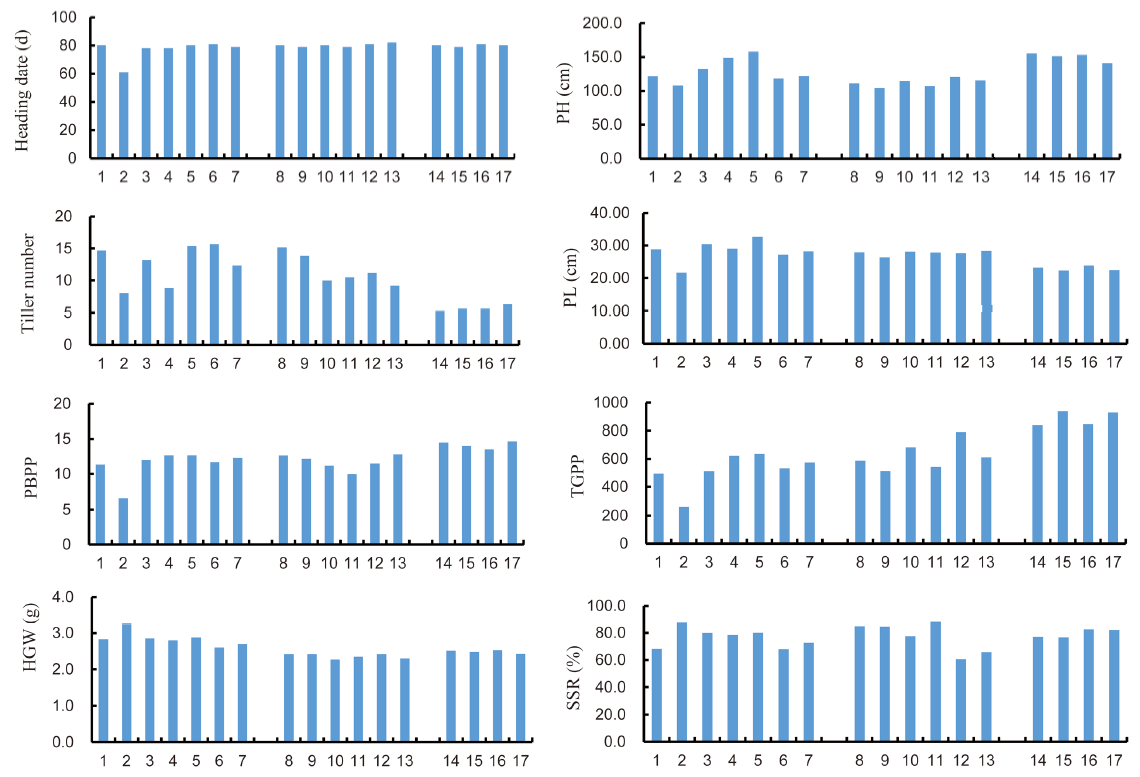
Fig. 2. Morphometric data on agronomic performance of Wonder Rice Initiative for Food Security and Health (WISH) lines in different backgrounds. 1, IR63307; 2, Sensho; 3, WISH110:1-1-11-12-1; 4, WISH110:1-1-11-5-4; 5, WISH110:2-5-2-1-1; 6, WISH110:2-5-2-12-1; 7, WISH110:2-5-2-22-1; 8, IRBB; 9, WISH48:1-3-1-1; 10, WISH48:1-3-2-1; 11, WISH48:1-3-18-1; 12, WISH48:1-3-20-1; 13, WISH48:1-3-21-1; 14, KP; 15, WISH40:1-3-16-11; 16, WISH40:1-3-3-9; 17, WISH40:1-3-7-7; PH, Plant height; PL, Panicle length; PBPP, Number of primary branches per panicle; TGPP, Total grain number per panicle; HGW, 100-grain weight; SSR, Seed-setting rate.
| Line name | Raw grain length (mm) | Raw grain width (mm) | Ratio of | Cooked grain length (mm) | Cooked grain width (mm) | Ratio of cooked grain length and width | Chalkiness (%) | Amylose content | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| grain length and width | ||||||||||||||||
| DS | WS | DS | WS | DS | WS | DS | WS | DS | WS | DS | WS | DS | WS | DS | WS | |
| Sensho | 6 | 6.1 | 2.9 | 2.8 | 2.1 | 2.2 | 10 | 9.9 | 4.3 | 4.4 | 2.4 | 2.3 | 6.8 | 16.4 | 21 | 17.4 |
| IR63307 | 5.8 | 5.9 | 2.8 | 2.7 | 2.1 | 2.2 | 9.2 | 8.9 | 4 | 3.9 | 2.3 | 2.3 | 7.3*a | 10.5*a | 24* | 21.1* |
| WISH 110:1-1-11-12-1 | 5.9 | 6 | 2.8 | 2.8 | 2.1 | 2.1 | 9.2 | 9.1 | 4.1 | 3.9 | 2.2 | 2.3 | 3.5*b | 8.3*b | 23* | 20.5* |
| WISH 110:1-1-11-5-4 | 5.9 | 5.9 | 2.8 | 2.7 | 2.1 | 2.2 | 9.5 | 9.4 | 4 | 3.9 | 2.4 | 2.4 | 1.6*c | 6.2*c | 23* | 22.4* |
| WISH 110:2-5-2-1-1 | 6.1 | 6 | 2.8 | 2.7 | 2.2 | 2.2 | 10 | 9.7 | 4.1 | 3.9 | 2.5 | 2.5 | 0.1*d | 3.2*d | 23* | 22.7* |
| WISH 110:2-5-2-12-1 | 5.7 | 5.6 | 2.8 | 2.7 | 2 | 2.1 | 9.1 | 8.8 | 4.1 | 3.8 | 2.2 | 2.3 | 1.7*c | 4.7*e | 23* | 21.6* |
| WISH 110:2-5-2-22-1 | 5.5 | 5.5 | 2.8 | 2.8 | 2 | 2 | 9.3 | 8.9 | 4.2 | 4 | 2.2 | 2.2 | 5.4*e | 6.9*f | 23* | 22.0* |
| IRBB | 6.9 | 6.8 | 2.2 | 2 | 3.1 | 3.4 | 10 | 10.3 | 4 | 3.5 | 2.6 | 2.9 | 0.3a | 0.4a | 13*a | 12.9*a |
| WISH 48:1-3-1-1 | 6.6 | 6.6 | 2.3 | 2.1 | 2.9 | 3.1 | 10 | 10.2 | 3.5 | 3.5 | 2.9 | 2.9 | 0.3a | 0.8b | 19*b | 15.3*b |
| WISH 48:1-3-18-1 | 6.4 | 6.8 | 2.3 | 2.1 | 2.8 | 3.2 | 11 | 10.5 | 3.5 | 3.2 | 3 | 3.3 | 1.2b | 0.2c | 21*c | 18.2*c |
| WISH 48:1-3-2-1 | 6.3 | 6.5 | 2.3 | 2.1 | 2.7 | 3.1 | 9.9 | 10.1 | 3.6 | 3.4 | 2.8 | 3 | 1.6c | 4.5d | 21*c | 19.6*d |
| WISH 48:1-3-20-1 | 6.7 | 6.9 | 2.3 | 2.1 | 2.9 | 3.3 | 9.9 | 10.4 | 3.6 | 3.5 | 2.8 | 3 | 5.6d | 7.9e | 20*d | 17.0*e |
| WISH 48:1-3-21-1 | 6.5 | 6.7 | 2.3 | 2.1 | 2.8 | 3.2 | 9.8 | 9.9 | 3.5 | 3.4 | 2.8 | 2.9 | 0.6e | 2.0f | 18*b | 18.9*f |
| KP | 5.1 | 5 | 2.9 | 2.7 | 1.8 | 1.9 | 8.7 | 8 | 4 | 3.8 | 2.2 | 2.1 | 7.5a | 8.5a | 23* | 21.3* |
| WISH 40:1-3-16-11 | 5 | 5.2 | 2.9 | 2.9 | 1.7 | 1.8 | 8.5 | 8.7 | 3.8 | 3.9 | 2.2 | 2.2 | 17.0b | 8.4a | 23* | 21.2* |
| WISH 40:1-3-3-9 | 5 | 5 | 2.9 | 2.8 | 1.7 | 1.8 | 8.4 | 8.3 | 3.9 | 3.8 | 2.2 | 2.2 | 10.2c | 14.5b | 24* | 22.0* |
| WISH 40:1-3-7-7 | 5.1 | 5 | 3 | 2.8 | 1.7 | 1.8 | 9.5 | 8.7 | 4.2 | 4 | 2.3 | 2.2 | 6.60d | 13.1c | 22* | 21.6* |
Supplemental Table 3. Grain quality of WISH lines having the pi21 allele for leaf blast resistance from Sensho measured during the dry (DS) and wet season (WS) of 2017.
| Line name | Raw grain length (mm) | Raw grain width (mm) | Ratio of | Cooked grain length (mm) | Cooked grain width (mm) | Ratio of cooked grain length and width | Chalkiness (%) | Amylose content | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| grain length and width | ||||||||||||||||
| DS | WS | DS | WS | DS | WS | DS | WS | DS | WS | DS | WS | DS | WS | DS | WS | |
| Sensho | 6 | 6.1 | 2.9 | 2.8 | 2.1 | 2.2 | 10 | 9.9 | 4.3 | 4.4 | 2.4 | 2.3 | 6.8 | 16.4 | 21 | 17.4 |
| IR63307 | 5.8 | 5.9 | 2.8 | 2.7 | 2.1 | 2.2 | 9.2 | 8.9 | 4 | 3.9 | 2.3 | 2.3 | 7.3*a | 10.5*a | 24* | 21.1* |
| WISH 110:1-1-11-12-1 | 5.9 | 6 | 2.8 | 2.8 | 2.1 | 2.1 | 9.2 | 9.1 | 4.1 | 3.9 | 2.2 | 2.3 | 3.5*b | 8.3*b | 23* | 20.5* |
| WISH 110:1-1-11-5-4 | 5.9 | 5.9 | 2.8 | 2.7 | 2.1 | 2.2 | 9.5 | 9.4 | 4 | 3.9 | 2.4 | 2.4 | 1.6*c | 6.2*c | 23* | 22.4* |
| WISH 110:2-5-2-1-1 | 6.1 | 6 | 2.8 | 2.7 | 2.2 | 2.2 | 10 | 9.7 | 4.1 | 3.9 | 2.5 | 2.5 | 0.1*d | 3.2*d | 23* | 22.7* |
| WISH 110:2-5-2-12-1 | 5.7 | 5.6 | 2.8 | 2.7 | 2 | 2.1 | 9.1 | 8.8 | 4.1 | 3.8 | 2.2 | 2.3 | 1.7*c | 4.7*e | 23* | 21.6* |
| WISH 110:2-5-2-22-1 | 5.5 | 5.5 | 2.8 | 2.8 | 2 | 2 | 9.3 | 8.9 | 4.2 | 4 | 2.2 | 2.2 | 5.4*e | 6.9*f | 23* | 22.0* |
| IRBB | 6.9 | 6.8 | 2.2 | 2 | 3.1 | 3.4 | 10 | 10.3 | 4 | 3.5 | 2.6 | 2.9 | 0.3a | 0.4a | 13*a | 12.9*a |
| WISH 48:1-3-1-1 | 6.6 | 6.6 | 2.3 | 2.1 | 2.9 | 3.1 | 10 | 10.2 | 3.5 | 3.5 | 2.9 | 2.9 | 0.3a | 0.8b | 19*b | 15.3*b |
| WISH 48:1-3-18-1 | 6.4 | 6.8 | 2.3 | 2.1 | 2.8 | 3.2 | 11 | 10.5 | 3.5 | 3.2 | 3 | 3.3 | 1.2b | 0.2c | 21*c | 18.2*c |
| WISH 48:1-3-2-1 | 6.3 | 6.5 | 2.3 | 2.1 | 2.7 | 3.1 | 9.9 | 10.1 | 3.6 | 3.4 | 2.8 | 3 | 1.6c | 4.5d | 21*c | 19.6*d |
| WISH 48:1-3-20-1 | 6.7 | 6.9 | 2.3 | 2.1 | 2.9 | 3.3 | 9.9 | 10.4 | 3.6 | 3.5 | 2.8 | 3 | 5.6d | 7.9e | 20*d | 17.0*e |
| WISH 48:1-3-21-1 | 6.5 | 6.7 | 2.3 | 2.1 | 2.8 | 3.2 | 9.8 | 9.9 | 3.5 | 3.4 | 2.8 | 2.9 | 0.6e | 2.0f | 18*b | 18.9*f |
| KP | 5.1 | 5 | 2.9 | 2.7 | 1.8 | 1.9 | 8.7 | 8 | 4 | 3.8 | 2.2 | 2.1 | 7.5a | 8.5a | 23* | 21.3* |
| WISH 40:1-3-16-11 | 5 | 5.2 | 2.9 | 2.9 | 1.7 | 1.8 | 8.5 | 8.7 | 3.8 | 3.9 | 2.2 | 2.2 | 17.0b | 8.4a | 23* | 21.2* |
| WISH 40:1-3-3-9 | 5 | 5 | 2.9 | 2.8 | 1.7 | 1.8 | 8.4 | 8.3 | 3.9 | 3.8 | 2.2 | 2.2 | 10.2c | 14.5b | 24* | 22.0* |
| WISH 40:1-3-7-7 | 5.1 | 5 | 3 | 2.8 | 1.7 | 1.8 | 9.5 | 8.7 | 4.2 | 4 | 2.3 | 2.2 | 6.60d | 13.1c | 22* | 21.6* |
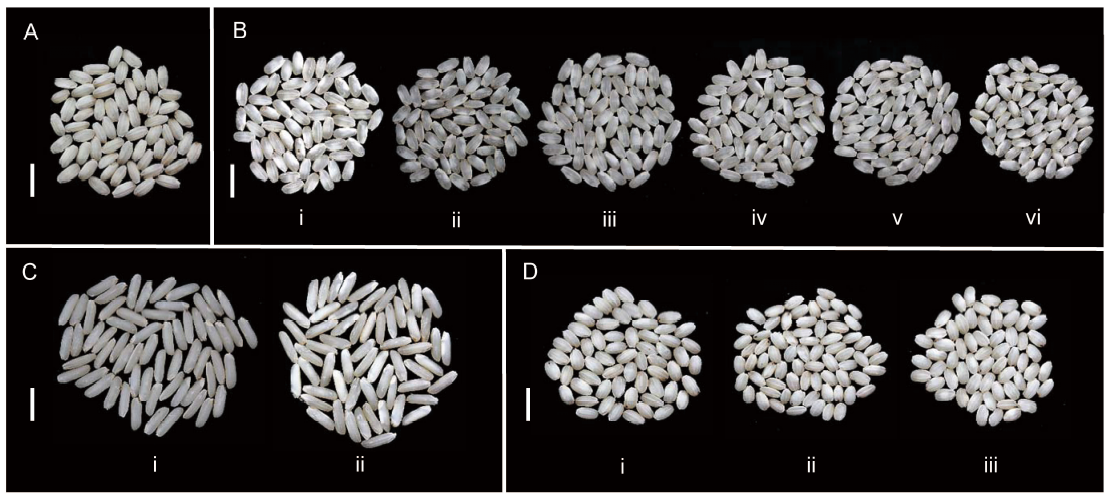
Fig. 3. Variation in the overall chalkiness degree of Wonder Rice Initiative for Food Security and Health (WISH) lines in different genetic backgrounds. A, Grains of Sensho. B, Grains of IR63307 (i), WISH110:2-5-2-1-1 (ii), WISH110:1-1-11-5-4 (iii), WISH110:2-5-2-12-1 (iv), WISH110:1-1-11-12-1 (v) and WISH110:2-5-2-22-1 (vi) in the background of IR63307, showing significantly lower chalkiness degrees compared to the recurrent parent. C, Grains of IRBB (i) and WISH48:1-3-20-2 (ii) in the IRBB background, showing a significantly higher chalkiness degree compared to the recurrent parent. D, Grains of KP (i), WISH40:1-3-3-9 (ii) and WISH40:1-3-16-11 (iii) in the KP background, showing higher chalkiness degrees compared to the recurrent parent. Bars = 10 mm.
| [1] | Bonman J M, Vergel De Dios T I, Khin M M. 1986. Physiological specialization of Pyricularia oryzae in the Philippines. Plant Dis, 70: 767-769. |
| [2] | Chen H, Iqbal M, Yang R C, Spanner D. 2016. Effect of Lr34/Yr18 on agronomic and quality traits in spring wheat mapping population and implications for breeding. Mol Breeding, 36: 53. |
| [3] | Cherif M, Harrabi M. 1993. Transgressive segregation for resistance to Pyrenophora teres in barley. Plant Pathol, 42(4): 617-621. |
| [4] | Dean R A, Talbot N J, Ebbole D J, Farman M L, Mitchell T K, Orbach M J, Thon M, Kulkarni R, Xu J R, Pan H Q, Read N D, Lee Y H, Carbone I, Brown D, Oh Y Y, Donofrio N, Jeong J S, Soanes D M, Djonovic S, Kolomiets E, Rehmeyer C, Li W X, Harding M, Kim S, Lebrun M H, Bohnert H, Coughlan S, Butler J, Calvo S, Ma L J, Nicol R, Purcell S, Nusbaum C, Galagan J E, Birren B W. 2005. The genome, sequence of the rice blast fungus Magnaporthe grisea. Nature, 434: 980-986. |
| [5] | de Vicente M C, Tanksley S D. 1993. QTL analysis of transgressive segregation in an interspecific tomato cross. Genetics, 134: 585-596. |
| [6] | Fujita D, Ebron L A, Kobayashi N, Fukuta Y. 2009. DNA marker analysis of blast resistance Pib and Pita in IRRI-bred rice varieties comparing with gene estimation by a differential system. In: Wang G L, Valent B. Advances in Genetics, Genomics and Control of Rice. Netherlands: Springer: 315-324. |
| [7] | Fukuoka S, Okuno K. 2001. QTL analysis and mapping of pi21, a recessive gene for field resistance to rice blast in Japanese upland rice. Theor Appl Genet, 103: 185-190. |
| [8] | Fukuoka S, Saka N, Koga H, Ono K, Shimizu T, Ebaba K, Hayashi N, Takahashi A, Hirochika H, Okuno K, Yano M. 2009. Loss of function of a proline-containing protein confers durable disease resistance in rice. Science, 325: 998-1001. |
| [9] | Fukuoka S, Mizobuchi R, Saka N, Ivan S, Matsumoto T, Okuno K, Yano M. 2012. A multiple gene complex on rice chromosome 4 is involved in durable resistance to rice blast. Theor Appl Genet, 125(3): 551-599. |
| [10] | Fukuoka S, Saka N, Mizukami Y, Koga H, Yamanouchi U, Yoshioka Y, Hayashi N, Ebana K, Mizobuchi R, Yano M. 2015. Gene pyramiding enhances durable blast disease resistance in rice. Sci Rep, 5: 7773. |
| [11] | Fukuta Y, Teleblanco-Yanoria M J, Imbe T, Tsunematsu H, Kato H, Ban T, Ebron L A, Hayashi N, Ando I, Khush G S. 2004. Monogenic lines as an international standard differential set for blast resistance in rice (Oryza sativa L.). Rice Genet Newsl, 21: 70. |
| [12] | Goto I. 1970. Genetic studies on the resistance of rice plant to the blast fungus: I. Inheritance of resistance in crosses Sensho × H-79 and Imochishirazu × H-79. Ann Phytopathol Soc Jpn, 36: 304-312. |
| [13] | Hayashi N, Kobayashi N, Vera Cruz C M, Fukuta Y. 2009. Protocols for sampling of disease specimens and evaluation of blast disease in rice Japan International Research Center for Agricultural Sciences (JIRCAS), Tsukuba, Japan. Work Rep, 6: 17-33. |
| [14] | Horo J T, Fuji T, Yamashita Y, McGoey S, Koizumi S. 2016. Rice blast control efficacy of three genes ( Pib, pi21 and Pb1) conferring complete and partial resistance. Jpn Agric Res Q, 50(3): 209-217. |
| [15] | Huang N, Angeles E R, Domingo J, Magpantay G, Singh S, Zhang G, Kumaravadivel N, Bennet J, Khush G S. 1997. Pyramiding of bacterial blight resistance genes in rice: Marker-assisted selection using RFLP and PCR. Theor Appl Genet, 96(3): 313-320. |
| [16] | Imbe T, Oba S, Yanoria M J T, Tsunematsu H. 1997. A new gene for blast resistance in rice cultivar, IR24. Rice Genet Newsl, 14: 60-61. |
| [17] | Imbe T, Tsunematsu H, Kato H, Khush G S. 1998. Genetic analysis of blast resistance in IR varieties and resistant breeding strategy. In: Tharreau D, Lebrun M H, Talbot N J, Notteghem J L. Advances in Rice Blast Research: Proceedings of the 2nd International Rice Blast Conference. Montpellier, France: 1-8. |
| [18] | International Standardization Organization. 2007. Rice: Determination of Amylose Content. Part 2: Routine Methods (ISO6647-2). International Standardization Organization. |
| [19] | IRRI (International Rice Research Institute). 2013. Statistical Tool for Agricultural Research (STAR): Plant Breeding, Genetics, and Biotechnology. Los Baños, the Philippines: IRRI. |
| [20] | IRRI (International Rice Research Institute). 2014. Standard Evaluation System for Rice (SES). 5th edn. Los Baños, the Philippines: IRRI: 57. |
| [21] | Jeung J U, Kim B R, Cho Y C, Han S S, Moon H P, Lee Y T, Jena K K. 2007. A novel gene Pi40(t), linked to the DNA markers derived from NBS-LRR motifs confers broad spectrum of blast resistance in rice. Theor Appl Genet, 115: 1163-1177. |
| [22] | Kato T, Endo I, Yano M, Sasaki T, Inoue M, Kudo S. 2002. Mapping of quantitative trait loci for field resistance to rice blast in upland rice, ‘Sensho’. Breeding Res, 4(3): 119-124. |
| [23] | Kawasaki-Tanaka A, Fukuta Y. 2014. Genetic variation in resistance to blast disease (Pyricularia oryzae Cavara) in Japanese rice(Oryza sativa L.), as determined using a differential system. Breeding Sci, 64(2): 183-192. |
| [24] | Kiyosawa S, Ling Z. 2001. Genetic studies on rice blast relationships. In: Sreenivasaprasad S, Johnson R. Major Fungal Disease in Rice. Dordrecht: Kluwer Academic: 145-162. |
| [25] | Kou Y J, Wang S P. 2012. Toward an understanding of the molecular basis of quantitative disease resistance in rice. J Biotechnol, 159(4): 283-290. |
| [26] | Li Y C, Yuan L P. 1985. Genetic analysis of fertility restoration in male sterile lines of rice. In: Banta S J. Rice Genetics. Manila, the Philippines: International Rice Research Institute: 617-632. |
| [27] | Miura K, Agetsuma M, Kitano H, Yoshimura A, Matsuoka M, Jacobsen S E, Ashikari M. 2009. A metastable DWARF1 epigenetic mutant affecting plant stature in rice. Proc Natl Acad Sci USA, 106: 11218-11223. |
| [28] | Piffanelli P, Zhou F, Casais C, Orme J, Jarosch B, Schaffrath U, Collins N C, Panstruga R, Schulze-Lefert P. 2002. The barley MLO modulator of defense and cell death is responsive to biotic and abiotic stress stimuli. Plant Physiol, 129(3): 1076-1085. |
| [29] | Prahalada G D, Ramkumar G, Hechanova S L, Vinarao R, Jena K K. 2017. Exploring key blast and bacterial blight resistance genes in genetically diverse rice accessions through molecular and phenotypic evaluation. Crop Sci, 57: 1881-1892. |
| [30] | Ram S, Mishra B. 2010. Cereals: Processing and Nutritional Quality. New Delhi, India: New India Publishing Agency: 326. |
| [31] | Rick C M, Smith P G. 1953. Novel variation in tomato species hybrids. Am Nat, 87: 359-375. |
| [32] | Riesberg L H, Archer M A, Wayne R K. 1999. Transgressive segregation, adaptation and speciation. Heredity, 83: 363-372. |
| [33] | Sallaud C, Lorieux M, Roumen E, Tharreau D, Berruyer R, Svestasrani P, Garsmeur O, Ghesquiere A, Notteghem J L. 2003. Identification of five new blast resistance genes in the highly blast-resistant rice variety IR64 using a QTL mapping strategy. Theor Appl Genet, 106: 794-803. |
| [34] | Scheuermann K K, Raimondi J V, Marschalek R, Andrade A, Wickert E. 2012. Magnaporthe oryzae genetic diversity and its outcomes on the search for durable resistance. In: Caliskan M. The Molecular Basis of Plant Genetic Diversity. Intech Open Access Publisher: 331-356. |
| [35] | Selisana S M, Yanoria M J, Quime B, Chaipanya C, Lu G, Opulencia R, Wang G L, Mitchel T, Correll J, Talbot N J, Leung H, Zhou B. 2017. Avirulence (AVR) gene-based diagnosis complements existing pathogen surveillance tools for effective deployment of resistance (R) genes against rice blast disease. Phytopathology, 107(6): 711-720. |
| [36] | Séré Y, Onasanya A, Afolabi A, Mignouna H D, Akator K. 2007. Genetic diversity of the blast fungus Magnaporthe grisea(Hebert) Barr, in Burkina Faso. Afr J Biotechnol, 6: 2568-2577. |
| [37] | Sharma T R, Rai A K, Gupta S K, Vijayan J, Devanna B N, Ray S. 2012. Rice blast management through host-plant resistance: Retrospect and prospects. Agric Res, 1: 37-52. |
| [38] | Shim J, Torollo G, Angeles-Shim R B, Cabunagan R C, Choi I R, Yeo U S, Ha W G. 2015. Rice tungro spherical virus resistance into photoperiod-insensitive japonica rice by marker-assisted selection. Breeding Sci, 65(4): 345-351. |
| [39] | Suh J P, Roh J H, Cho Y C, Han S S, Kim Y G, Jena K K. 2009. The Pi40 gene for durable resistance to rice blast and molecular analysis of Pi40-advanced backcross breeding lines. Phytopathology, 99(3): 243-250. |
| [40] | Teleblanco-Yanoria M J, Imber T, Kato H, Tsunematsu H, Ebron L A, Vera Cruz C M, Kobayashi N, Fukuta Y. 2008. A set of standard differential blast isolates (Magnaporthe grisea (Hebert) Barr.) from the Philippines for rice (Oryza sativa L.) resistance. J Agric Res Quart, 42(1): 23-34. |
| [41] | Teleblanco-Yanoria M J, Koide Y, Fukuta Y, Imbe T, Tsunematsu H, Kato H, Ebron L A, Nguyen T M N, Kobayashi N. 2011. A set of differential lines of indica-type rice variety CO39 as differential varieties for blast resistance. Mol Breeding, 27(3): 357-373. |
| [42] | Uga Y, Okuno K, Yano M. 2011. Dro1, a major QTL involved in deep rooting of rice under upland field conditions. J Exp Bot, 62(8): 2485-2494. |
| [43] | Uga Y, Yamamoto E, Kanno N, Kawai S, Mizubayashi T, Fukuoka S. 2013. A major QTL controlling deep rooting on rice chromosome 4. Sci Rep, 3: 3040. |
| [44] | Vasudevan K, Vera Cruz C M, Gruissem W, Bhullar N K. 2014. Large scale germplasm screening for identification of novel blast resistance sources. Front Plant Sci, 5: 505. |
| [45] | Vega U, Frey K J. 1980. Transgressive segregation in inter- and intra-specific crosses of barley. Euphytica, 29(3): 585-594. |
| [46] | Wallwork H, Johnson R. 1984. Transgressive segregation for resistance to yellow rust in wheat. Euphytica, 33(1): 123-132. |
| [47] | Wang C L, Ulloa M, Mullens T R, Yu J Z, Roberts P A. 2012. QTL analysis for transgressive resistance to root-knot nematode in interspecific cotton (Gossypium spp) progeny derived from susceptible parents. PLoS One, 7: e344874. |
| [48] | Yasuda N, Mitsunaga T, Hayashi K, Koizumi S, Fujita Y. 2015. Effects of pyramiding quantitative resistance genes pi21, Pi34 and Pi35 on rice leaf blast disease. Plant Dis, 99(7): 904-909. |
| [49] | Yue B, Xue W Y, Xiong L Z, Yu X Q, Luo L J, Cui K H, Jin D M, Xing Y Z, Zhang Q F. 2006. Genetic basis of drought resistance at reproductive stage in rice: Separation of drought tolerance from drought avoidance. Genetics, 172(2): 1213-1228. |
| [50] | Zeigler R S, Tohme J, Nelson R, Levy M, Corre F J. 1994. Lineage exclusion: A proposal for linking blast population analysis to breeding. In: Zeigler R S, Leong S A, Teeng P S. Rice Blast Disease. Wallingford: CAB International: 267-292. |
| [51] | Zeng Y X, Xia L Z, Wen Z H, Ji Z J, Zeng D L, Qian Q, Yang C D. 2015. Mapping resistant QTLs for rice sheath blight disease with a doubled haploid population. J Integr Agric, 14(5): 801-810. |
| [1] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [2] | Wu Zhongling, Qiu Jiehua, Shi Huanbin, Lin Chuyu, Yue Jiangnan, Liu Zhiquan, Xie Wei, Naweed I. Naqvi, Kou Yanjun, Tao Zeng. Polycomb Repressive Complex 2-Mediated H3K27 Trimethylation Is Required for Pathogenicity in Magnaporthe oryzae [J]. Rice Science, 2022, 29(4): 363-374. |
| [3] | Suhas Gorakh Karkute, Vishesh Kumar, Mohd Tasleem, Dwijesh Chandra Mishra, Krishna Kumar Chaturvedi, Anil Rai, Amitha Mithra Sevanthi, Kishor Gaikwad, Tilak Raj Sharma, Amolkumar U. Solanke. Genome-Wide Analysis of von Willebrand Factor A Gene Family in Rice for Its Role in Imparting Biotic Stress Resistance with Emphasis on Rice Blast Disease [J]. Rice Science, 2022, 29(4): 375-384. |
| [4] | Zhou Ying, Wan Tao, Yuan Bin, Lei Fang, Chen Meijuan, Wang Qiong, Huang Ping, Kou Shuyan, Qiu Wenxiu, Liu Li. Improving Rice Blast Resistance by Mining Broad-Spectrum Resistance Genes at Pik Locus [J]. Rice Science, 2022, 29(2): 133-142. |
| [5] | R. Abdul Fiyaz, D. Shivani, K. Chaithanya, K. Mounika, M. Chiranjeevi, G. S. Laha, B. C. Viraktamath, L. V. Subba Rao, R. M. Sundaram. Genetic Improvement of Rice for Bacterial Blight Resistance: Present Status and Future Prospects [J]. Rice Science, 2022, 29(2): 118-132. |
| [6] | Muduli Lakesh, Kumar Pradhan Sukanta, Mishra Abinash, Nath Bastia Debendra, Chandra Samal Kailash, Kumar Agrawal Pawan, Dash Manasi. Understanding Brown Planthopper Resistance in Rice: Genetics, Biochemical and Molecular Breeding Approaches [J]. Rice Science, 2021, 28(6): 532-546. |
| [7] | Yong Yang, Qiujun Lin, Xinyu Chen, Weifang Liang, Yuwen Fu, Zhengjin Xu, Yuanhua Wu, Xuming Wang, Jie Zhou, Chulang Yu, Chengqi Yan, Qiong Mei, Jianping Chen. Characterization and Proteomic Analysis of Novel Rice Lesion Mimic Mutant with Enhanced Disease Resistance [J]. Rice Science, 2021, 28(5): 466-478. |
| [8] | Saichompoo Uthomphon, Narumol Possawat, Nakwilai Pawat, Thongyos Peeranut, Nanta Aekchupong, Tippunya Patompong, Ruengphayak Siriphat, Itthisoponkul Teerarat, Bueraheng Niranee, Cheabu Sulaiman, Malumpong Chanate. Breeding Novel Short Grain Rice for Tropical Region to Combine Important Agronomical Traits, Biotic Stress Resistance and Cooking Quality in Koshihikari Background [J]. Rice Science, 2021, 28(5): 479-792. |
| [9] | Mishra Rukmini, Zheng Wei, Kumar Joshi Raj, Kaijun Zhao. Genome Editing Strategies Towards Enhancement of Rice Disease Resistance [J]. Rice Science, 2021, 28(2): 133-145. |
| [10] | Junhua Lu, Xuemei Yang, Jinfeng Chen, Tingting Li, Zijin Hu, Ying Xie, Jinlu Li, Jiqun Zhao, Mei Pu, Hui Feng, Jing Fan, Yanyan Huang, Jiwei Zhang, Wenming Wang, Yan Li. Osa-miR439 Negatively Regulates Rice Immunity Against Magnaporthe oryzae [J]. Rice Science, 2021, 28(2): 156-165. |
| [11] | Yanchang Luo, Tingchen Ma, Teo Joanne, Zhixiang Luo, Zefu Li, Jianbo Yang, Zhongchao Yin. Marker-Assisted Breeding of Thermo-Sensitive Genic Male Sterile Line 1892S for Disease Resistance and Submergence Tolerance [J]. Rice Science, 2021, 28(1): 89-98. |
| [12] | Ning Xiao, Yunyu Wu, Aihong Li. Strategy for Use of Rice Blast Resistance Genes in Rice Molecular Breeding [J]. Rice Science, 2020, 27(4): 263-277. |
| [13] | Ting Chen, Zheng Chen, Prakash Sathe Atul, Zhihong Zhang, Liangjian Li, Huihui Shang, Shaoqing Tang, Xiaobo Zhang, Jianli Wu. Characterization of a Novel Gain-of-Function Spotted-Leaf Mutant with Enhanced Disease Resistance in Rice [J]. Rice Science, 2019, 26(6): 372-383. |
| [14] | Jiehua Qiu, Shuai Meng, Yizhen Deng, Shiwen Huang, Yanjun Kou. Ustilaginoidea virens: A Fungus Infects Rice Flower and Threats World Rice Production [J]. Rice Science, 2019, 26(4): 199-206. |
| [15] | Tao Chen, Hao Wu, Ya-dong Zhang, Zhen Zhu, Qi-yong Zhao, Li-hui Zhou, Shu Yao, Ling Zhao, Xin Yu, Chun-fang Zhao, Cai-lin Wang. Genetic Improvement of Japonica Rice Variety Wuyujing 3 for Stripe Disease Resistance and Eating Quality by Pyramiding Stv-bi and Wx-mq [J]. Rice Science, 2016, 23(2): 69-77. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||