
Rice Science ›› 2015, Vol. 22 ›› Issue (4): 180-188.DOI: 10.1016/S1672-6308(14)60293-6
• Orginal Article • Previous Articles Next Articles
Mao-bai Li1,2, Hui Wang1, Li-ming Cao1( )
)
Received:2014-09-23
Accepted:2015-02-11
Online:2015-07-28
Published:2015-05-27
About author:Corresponding author:CAO Li-ming (clm079@163.com)
Mao-bai Li, Hui Wang, Li-ming Cao. Evaluation of Population Structure, Genetic Diversity and Origin of Northeast Asia Weedy Rice Based on Simple Sequence Repeat Markers[J]. Rice Science, 2015, 22(4): 180-188.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.ricesci.org/EN/10.1016/S1672-6308(14)60293-6
| Origin | Code for each material a | Year |
| Qingpu, Shanghai, China | Qp-1, Qp-2, Qp-3, Qp-4, Qp-5 | 2008 |
| Jiading, Shanghai, China | Jd-1, Jd-2, Jd-3, Jd-4, Jd-5, Jd-6 | 2007 |
| Nanhui, Shanghai, China | Nh-1, Nh-2, Nh-3, Nh-4 | 2010 |
| Pudong, Shanghai, China | Pd-1, Pd-2, Pd-3, Pd-4, Pd-5 | 2009 |
| Chongming, Shanghai, China | Cm-1, Cm-2, Cm-3, Cm-4, Cm-5 | 2008 |
| Fengxian, Shanghai, China | Fx-1, Fx-2, Fx-3, Fx-4, Fx-5 | 2009 |
| Taizhou, Jiangsu, China | Tz-1, Tz-2, Tz-3, Tz-4, Tz-5 | 2005 |
| Shenyang, Liaoning, China | Sy-1, Sy-2, Sy-3, Sy-4, Sy-5 | 2000 |
| Tieling, Liaoning, China | Tl-1, Tl-2, Tl-3, Tl-4, Tl-5 | 2004 |
| Haicheng, Liaoning, China | Hc-1, Hc-2, Hc-3, Hc-4, Hc-5 | 2003 |
| Dandong, Liaoning, China | Dd-1, Dd-2, Dd-3, Dd-4, Dd-5 | 2004 |
| Gongzhuling, Jilin, China | Gz-1, Gz-2, Gz-3, Gz-4, Gz-5 | 2006 |
| Jiamusi, Heilongjiang, China | Jm-1, Jm-2, Jm-3, Jm-4, Jm-5 | 2005 |
| Yesan, Chungcheong, Korea | Ys-1, Ys-2, Ys-3, Ys-4, Ys-5 | 2000 |
| Seosan, Chungcheong, Korea | Ss-1, Ss-2, Ss-3, Ss-4, Ss-5, Ss-6 | 2000 |
| Qp, Qingpu; Jd, Jiading; Nh, Nanhui; Pd, Pudong; Cm, Chongming; Fx, Fengxian; Tz, Taizhou; Sy, Shenyang; Tl, Tieling; Hc, Haicheng; Dd, Dandong; Gz, Gongzhuling; Jm, Jiamusi; Ys, Yesan; Ss, Seosan. a The numbers represent weedy rice accessions from each location. | ||
Table 1 Origins, codes and collected years of Northeast Asia weedy rice accessions used for SSR analysis.
| Origin | Code for each material a | Year |
| Qingpu, Shanghai, China | Qp-1, Qp-2, Qp-3, Qp-4, Qp-5 | 2008 |
| Jiading, Shanghai, China | Jd-1, Jd-2, Jd-3, Jd-4, Jd-5, Jd-6 | 2007 |
| Nanhui, Shanghai, China | Nh-1, Nh-2, Nh-3, Nh-4 | 2010 |
| Pudong, Shanghai, China | Pd-1, Pd-2, Pd-3, Pd-4, Pd-5 | 2009 |
| Chongming, Shanghai, China | Cm-1, Cm-2, Cm-3, Cm-4, Cm-5 | 2008 |
| Fengxian, Shanghai, China | Fx-1, Fx-2, Fx-3, Fx-4, Fx-5 | 2009 |
| Taizhou, Jiangsu, China | Tz-1, Tz-2, Tz-3, Tz-4, Tz-5 | 2005 |
| Shenyang, Liaoning, China | Sy-1, Sy-2, Sy-3, Sy-4, Sy-5 | 2000 |
| Tieling, Liaoning, China | Tl-1, Tl-2, Tl-3, Tl-4, Tl-5 | 2004 |
| Haicheng, Liaoning, China | Hc-1, Hc-2, Hc-3, Hc-4, Hc-5 | 2003 |
| Dandong, Liaoning, China | Dd-1, Dd-2, Dd-3, Dd-4, Dd-5 | 2004 |
| Gongzhuling, Jilin, China | Gz-1, Gz-2, Gz-3, Gz-4, Gz-5 | 2006 |
| Jiamusi, Heilongjiang, China | Jm-1, Jm-2, Jm-3, Jm-4, Jm-5 | 2005 |
| Yesan, Chungcheong, Korea | Ys-1, Ys-2, Ys-3, Ys-4, Ys-5 | 2000 |
| Seosan, Chungcheong, Korea | Ss-1, Ss-2, Ss-3, Ss-4, Ss-5, Ss-6 | 2000 |
| Qp, Qingpu; Jd, Jiading; Nh, Nanhui; Pd, Pudong; Cm, Chongming; Fx, Fengxian; Tz, Taizhou; Sy, Shenyang; Tl, Tieling; Hc, Haicheng; Dd, Dandong; Gz, Gongzhuling; Jm, Jiamusi; Ys, Yesan; Ss, Seosan. a The numbers represent weedy rice accessions from each location. | ||
| Origin | Code | Accession name |
| Shanghai, China | C-Sh | Jinfeng, Youfeng, Hanfeng, Qiufeng |
| Jiangsu, China | C-Js | Xiushui 7, Xiushui 9, Xiushui 61, Xiushui 110, Wuyunjing 7, Wuyunjing 3, Zhendao 88 |
| Liaoning, China | C-Ln | Liaojing 9, Liaojing 294, Liaojing 371, Liaoxing 15, Shendao 7 |
| Chungcheong, Korea | C-Cc | Ilpum, Singeumo, Gopum, Dongjin 2, Yeon 308 |
Table 2 List of cultivated rice collected in Northeast Asia used for SSR analysis.
| Origin | Code | Accession name |
| Shanghai, China | C-Sh | Jinfeng, Youfeng, Hanfeng, Qiufeng |
| Jiangsu, China | C-Js | Xiushui 7, Xiushui 9, Xiushui 61, Xiushui 110, Wuyunjing 7, Wuyunjing 3, Zhendao 88 |
| Liaoning, China | C-Ln | Liaojing 9, Liaojing 294, Liaojing 371, Liaoxing 15, Shendao 7 |
| Chungcheong, Korea | C-Cc | Ilpum, Singeumo, Gopum, Dongjin 2, Yeon 308 |
| Primer | Chr | SSR motif | Forward sequence | Reverse sequence | EPS | Na | Ne | I |
| RM1 | 1 | (GA)26 | GCGAAAACACAATGCAAAAA | GCGTTGGTTGGACCTGAC | 113 | 6 | 2.721 | 1.250 |
| RM9 | 1 | (GA)15GT(GA)2 | GGTGCCATTGTCGTCCTC | ACGGCCCTCATCACCTTC | 136 | 3 | 1.236 | 0.398 |
| RM14 | 1 | (GA)18 | CCGAGGAGAGGAGTTCGAC | GTGCCAATTTCCTCGAAAAA | 191 | 4 | 2.416 | 1.041 |
| RM23 | 1 | (GA)15 | CATTGGAGTGGAGGCTGG | GTCAGGCTTCTGCCATTCTC | 145 | 3 | 1.903 | 0.733 |
| RM212 | 1 | (CT)24 | CCACTTTCAGCTACTACCAG | CACCCATTTGTCTCTCATTATG | 136 | 2 | 1.403 | 0.462 |
| RM237 | 1 | (CT)18 | CAAATCCCGACTGCTGTCC | TGGGAAGAGAGCACTACAGC | 130 | 2 | 1.492 | 0.512 |
| RM110 | 2 | (GA)15 | TCGAAGCCATCCACCAACGAAG | TCCGTACGCCGACGAGGTCGAG | 156 | 2 | 1.280 | 0.377 |
| RM425 | 2 | (CGG)9 | CCAACGAAGATTCGAAGCTC | CAGCACCATGAAGTCGCC | 126 | 3 | 2.103 | 0.814 |
| RM525 | 2 | (AAG)12 | GGCCCGTCCAAGAAATATTG | CGGTGAGACAGAATCCTTACG | 131 | 4 | 3.034 | 1.179 |
| RM1178 | 2 | (AG)13 | CAGTGGGCGAGCATAGGAG | ATCCTTTTCTCCCTCTCTCG | 112 | 3 | 2.998 | 1.098 |
| RM156 | 3 | (CGG)8 | GCCGCACCCTCACTCCCTCCTC | TCTTGCCGGAGCGCTTGAGGTG | 160 | 3 | 2.090 | 0.879 |
| RM218 | 3 | (TC)24ACT5(GT)11 | TGGTCAAACCAAGGTCCTTC | GACATACATTCTACCCCCGG | 148 | 3 | 1.943 | 0.766 |
| RM489 | 3 | (ATA)8 | ACTTGAGACGATCGGACACC | TCACCCATGGATGTTGTCAG | 271 | 4 | 2.175 | 0.893 |
| RM8209 | 3 | (AG)15 | AGGAGAAGAGGAATCTTTGC | CGATCGAGAGCTACTATTGC | 188 | 3 | 1.043 | 0.117 |
| RM241 | 4 | (CT)31 | GAGCCAAATAAGATCGCTGA | TGCAAGCAGCAGATTTAGTG | 138 | 2 | 1.729 | 0.613 |
| RM470 | 4 | (CTT)14 | TCCTCATCGGCTTCTTCTTC | AGAACCCGTTCTACGTCACG | 83 | 4 | 3.406 | 1.297 |
| RM567 | 4 | (GA)21 | ATCAGGGAAATCCTGAAGGG | GGAAGGAGCAATCACCACTG | 261 | 1 | 1.000 | 0.000 |
| RM5473 | 4 | (TC)20 | ACACGGAGATCCGAGACACGAG | CGAGATTAACGTCGTCCTC | 105 | 5 | 4.117 | 1.493 |
| RM31 | 5 | (GA)15 | GATCACGATCCACTGGAGCT | AAGTCCATTACTCTCCTCCC | 140 | 6 | 2.737 | 1.266 |
| RM188 | 5 | (CA)8 | TCCGCCTCTCCTCTCGCTTCCC | GCAACGCACAACCGAACCGAGC | 210 | 3 | 2.283 | 0.924 |
| RM430 | 5 | (GA)25 | AAACAACGACGTCCCTGATC | GTGCCTCCGTGGTTATGAAC | 173 | 5 | 2.922 | 1.217 |
| RM193 | 6 | (GCT)5 | CGCCTCTTCTTCCTCGCCTCCG | CGGGTCCATCCCCCCTCTCCTC | 189 | 1 | 1.000 | 0.000 |
| RM276 | 6 | (AG)8A3(GA)33 | CTCAACGTTGACACCTCGTG | TCCTCCATCGAGCAGTATCA | 149 | 4 | 2.113 | 0.960 |
| RM402 | 6 | (ATA)7 | GAGCCATGGAAAGATGCATG | TCAGCTGGCCTATGACAATG | 133 | 3 | 1.321 | 0.487 |
| RM469 | 6 | (AG)15 | AGCTGAACAAGCCCTGAAAG | GACTTGGGCAGTGTGACATG | 105 | 2 | 1.465 | 0.498 |
| RM557 | 6 | (AC)13 | GTGGCGAGATCTATGTGGTG | GCTTTGTGTGTGTGTGTGTG | 211 | 1 | 1.000 | 0.000 |
| RM234 | 7 | (CT)25 | ACAGTATCCAAGGCCCTGG | CACGTGAGACAAAGACGGAG | 156 | 2 | 1.729 | 0.613 |
| RM346 | 7 | (CTT)18 | CGAGAGAGCCCATAACTACG | ACAAGACGACGAGGAGGGAC | 175 | 3 | 2.132 | 0.867 |
| RM429 | 7 | (TG)10 | TCCCTCCAGCAATGTCTTTC | CCTTCATCTTGCTTTCCACC | 159 | 3 | 1.876 | 0.744 |
| RM500 | 7 | (AAG)9 | GAGCTTGCCAGAGTGGAAAG | GTTACACCGAGAGCCAGCTC | 259 | 1 | 1.000 | 0.000 |
| RM256 | 8 | (CT)21 | GACAGGGAGTGATTGAAGGC | GTTGATTTCGCCAAGGGC | 127 | 2 | 1.021 | 0.058 |
| RM264 | 8 | GA)27 | GTTGCGTCCTACTGCTACTTC | GATCCGTGTCGATGATTAGC | 178 | 5 | 2.586 | 1.149 |
| RM281 | 8 | (GA)21 | ACCAAGCATCCAGTGACCAG | GTTCTTCATACAGTCCACATG | 138 | 3 | 2.073 | 0.799 |
| RM408 | 8 | (CT)13 | CAACGAGCTAACTTCCGTCC | ACTGCTACTTGGGTAGCTGACC | 128 | 3 | 2.014 | 0.783 |
| RM515 | 8 | (GA)11 | TAGGACGACCAAAGGGTGAG | TGGCCTGCTCTCTCTCTCTC | 211 | 4 | 2.419 | 1.005 |
| RM201 | 9 | (CT)17 | CTCGTTTATTACCTACAGTACC | CTACCTCCTTTCTAGACCGATA | 158 | 2 | 1.741 | 0.617 |
| RM242 | 9 | (CT)26 | GGCCAACGTGTGTATGTCTC | TATATGCCAAGACGGATGGG | 225 | 3 | 1.916 | 0.807 |
| RM257 | 9 | (CT)24 | CAGTTCCGAGCAAGAGTACTC | GGATCGGACGTGGCATATG | 147 | 4 | 3.504 | 1.314 |
| RM566 | 9 | (AG)15 | ACCCAACTACGATCAGCTCG | CTCCAGGAACACGCTCTTTC | 239 | 3 | 1.879 | 0.802 |
| RM216 | 10 | (CT)18 | GCATGGCCGATGGTAAAG | TGTATAAAACCACACGGCCA | 146 | 3 | 1.291 | 0.436 |
| RM228 | 10 | (CA)6(GA)36 | CTGGCCATTAGTCCTTGG | GCTTGCGGCTCTGCTTAC | 154 | 7 | 4.632 | 1.693 |
| RM484 | 10 | (AT)9 | TCTCCCTCCTCACCATTGTC | TGCTGCCCTCTCTCTCTCTC | 299 | 1 | 1.000 | 0.000 |
| RM224 | 11 | (AAG)8(AG)13 | ATCGATCGATCTTCACGAGG | TGCTATAAAAGGCATTCGGG | 157 | 4 | 2.390 | 1.070 |
| RM286 | 11 | (GA)16 | GGCTTCATCTTTGGCGAC | CCGGATTCACGAGATAAACTC | 110 | 4 | 2.484 | 1.066 |
| RM17 | 12 | (GA)21 | TGCCCTGTTATTTTCTTCTCTC | GGTGATCCTTTCCCATTTCA | 184 | 3 | 1.349 | 0.495 |
| RM101 | 12 | (CT)37 | GTGAATGGTCAAGTGACTTAGGTGGC | ACACAACATGTTCCCTCCCATGC | 324 | 3 | 2.563 | 1.001 |
| RM277 | 12 | (GA)11 | CGGTCAAATCATCACCTGAC | CAAGGCTTGCAAGGGAAG | 124 | 2 | 1.929 | 0.675 |
| RM1103 | 12 | (AG)12 | CAGCTGCTGCTACTACACCG | CTACTCCACGTCCATGCATG | 216 | 2 | 1.792 | 0.634 |
| Chr, Chromosome; EPS, Expected PCR product size; Na, Observed number of alleles; Ne, Effective number of alleles; I, Shannon’s information index. | ||||||||
Table 3 Repeat mofitis, sequences and genetic diversity of SSR primers.
| Primer | Chr | SSR motif | Forward sequence | Reverse sequence | EPS | Na | Ne | I |
| RM1 | 1 | (GA)26 | GCGAAAACACAATGCAAAAA | GCGTTGGTTGGACCTGAC | 113 | 6 | 2.721 | 1.250 |
| RM9 | 1 | (GA)15GT(GA)2 | GGTGCCATTGTCGTCCTC | ACGGCCCTCATCACCTTC | 136 | 3 | 1.236 | 0.398 |
| RM14 | 1 | (GA)18 | CCGAGGAGAGGAGTTCGAC | GTGCCAATTTCCTCGAAAAA | 191 | 4 | 2.416 | 1.041 |
| RM23 | 1 | (GA)15 | CATTGGAGTGGAGGCTGG | GTCAGGCTTCTGCCATTCTC | 145 | 3 | 1.903 | 0.733 |
| RM212 | 1 | (CT)24 | CCACTTTCAGCTACTACCAG | CACCCATTTGTCTCTCATTATG | 136 | 2 | 1.403 | 0.462 |
| RM237 | 1 | (CT)18 | CAAATCCCGACTGCTGTCC | TGGGAAGAGAGCACTACAGC | 130 | 2 | 1.492 | 0.512 |
| RM110 | 2 | (GA)15 | TCGAAGCCATCCACCAACGAAG | TCCGTACGCCGACGAGGTCGAG | 156 | 2 | 1.280 | 0.377 |
| RM425 | 2 | (CGG)9 | CCAACGAAGATTCGAAGCTC | CAGCACCATGAAGTCGCC | 126 | 3 | 2.103 | 0.814 |
| RM525 | 2 | (AAG)12 | GGCCCGTCCAAGAAATATTG | CGGTGAGACAGAATCCTTACG | 131 | 4 | 3.034 | 1.179 |
| RM1178 | 2 | (AG)13 | CAGTGGGCGAGCATAGGAG | ATCCTTTTCTCCCTCTCTCG | 112 | 3 | 2.998 | 1.098 |
| RM156 | 3 | (CGG)8 | GCCGCACCCTCACTCCCTCCTC | TCTTGCCGGAGCGCTTGAGGTG | 160 | 3 | 2.090 | 0.879 |
| RM218 | 3 | (TC)24ACT5(GT)11 | TGGTCAAACCAAGGTCCTTC | GACATACATTCTACCCCCGG | 148 | 3 | 1.943 | 0.766 |
| RM489 | 3 | (ATA)8 | ACTTGAGACGATCGGACACC | TCACCCATGGATGTTGTCAG | 271 | 4 | 2.175 | 0.893 |
| RM8209 | 3 | (AG)15 | AGGAGAAGAGGAATCTTTGC | CGATCGAGAGCTACTATTGC | 188 | 3 | 1.043 | 0.117 |
| RM241 | 4 | (CT)31 | GAGCCAAATAAGATCGCTGA | TGCAAGCAGCAGATTTAGTG | 138 | 2 | 1.729 | 0.613 |
| RM470 | 4 | (CTT)14 | TCCTCATCGGCTTCTTCTTC | AGAACCCGTTCTACGTCACG | 83 | 4 | 3.406 | 1.297 |
| RM567 | 4 | (GA)21 | ATCAGGGAAATCCTGAAGGG | GGAAGGAGCAATCACCACTG | 261 | 1 | 1.000 | 0.000 |
| RM5473 | 4 | (TC)20 | ACACGGAGATCCGAGACACGAG | CGAGATTAACGTCGTCCTC | 105 | 5 | 4.117 | 1.493 |
| RM31 | 5 | (GA)15 | GATCACGATCCACTGGAGCT | AAGTCCATTACTCTCCTCCC | 140 | 6 | 2.737 | 1.266 |
| RM188 | 5 | (CA)8 | TCCGCCTCTCCTCTCGCTTCCC | GCAACGCACAACCGAACCGAGC | 210 | 3 | 2.283 | 0.924 |
| RM430 | 5 | (GA)25 | AAACAACGACGTCCCTGATC | GTGCCTCCGTGGTTATGAAC | 173 | 5 | 2.922 | 1.217 |
| RM193 | 6 | (GCT)5 | CGCCTCTTCTTCCTCGCCTCCG | CGGGTCCATCCCCCCTCTCCTC | 189 | 1 | 1.000 | 0.000 |
| RM276 | 6 | (AG)8A3(GA)33 | CTCAACGTTGACACCTCGTG | TCCTCCATCGAGCAGTATCA | 149 | 4 | 2.113 | 0.960 |
| RM402 | 6 | (ATA)7 | GAGCCATGGAAAGATGCATG | TCAGCTGGCCTATGACAATG | 133 | 3 | 1.321 | 0.487 |
| RM469 | 6 | (AG)15 | AGCTGAACAAGCCCTGAAAG | GACTTGGGCAGTGTGACATG | 105 | 2 | 1.465 | 0.498 |
| RM557 | 6 | (AC)13 | GTGGCGAGATCTATGTGGTG | GCTTTGTGTGTGTGTGTGTG | 211 | 1 | 1.000 | 0.000 |
| RM234 | 7 | (CT)25 | ACAGTATCCAAGGCCCTGG | CACGTGAGACAAAGACGGAG | 156 | 2 | 1.729 | 0.613 |
| RM346 | 7 | (CTT)18 | CGAGAGAGCCCATAACTACG | ACAAGACGACGAGGAGGGAC | 175 | 3 | 2.132 | 0.867 |
| RM429 | 7 | (TG)10 | TCCCTCCAGCAATGTCTTTC | CCTTCATCTTGCTTTCCACC | 159 | 3 | 1.876 | 0.744 |
| RM500 | 7 | (AAG)9 | GAGCTTGCCAGAGTGGAAAG | GTTACACCGAGAGCCAGCTC | 259 | 1 | 1.000 | 0.000 |
| RM256 | 8 | (CT)21 | GACAGGGAGTGATTGAAGGC | GTTGATTTCGCCAAGGGC | 127 | 2 | 1.021 | 0.058 |
| RM264 | 8 | GA)27 | GTTGCGTCCTACTGCTACTTC | GATCCGTGTCGATGATTAGC | 178 | 5 | 2.586 | 1.149 |
| RM281 | 8 | (GA)21 | ACCAAGCATCCAGTGACCAG | GTTCTTCATACAGTCCACATG | 138 | 3 | 2.073 | 0.799 |
| RM408 | 8 | (CT)13 | CAACGAGCTAACTTCCGTCC | ACTGCTACTTGGGTAGCTGACC | 128 | 3 | 2.014 | 0.783 |
| RM515 | 8 | (GA)11 | TAGGACGACCAAAGGGTGAG | TGGCCTGCTCTCTCTCTCTC | 211 | 4 | 2.419 | 1.005 |
| RM201 | 9 | (CT)17 | CTCGTTTATTACCTACAGTACC | CTACCTCCTTTCTAGACCGATA | 158 | 2 | 1.741 | 0.617 |
| RM242 | 9 | (CT)26 | GGCCAACGTGTGTATGTCTC | TATATGCCAAGACGGATGGG | 225 | 3 | 1.916 | 0.807 |
| RM257 | 9 | (CT)24 | CAGTTCCGAGCAAGAGTACTC | GGATCGGACGTGGCATATG | 147 | 4 | 3.504 | 1.314 |
| RM566 | 9 | (AG)15 | ACCCAACTACGATCAGCTCG | CTCCAGGAACACGCTCTTTC | 239 | 3 | 1.879 | 0.802 |
| RM216 | 10 | (CT)18 | GCATGGCCGATGGTAAAG | TGTATAAAACCACACGGCCA | 146 | 3 | 1.291 | 0.436 |
| RM228 | 10 | (CA)6(GA)36 | CTGGCCATTAGTCCTTGG | GCTTGCGGCTCTGCTTAC | 154 | 7 | 4.632 | 1.693 |
| RM484 | 10 | (AT)9 | TCTCCCTCCTCACCATTGTC | TGCTGCCCTCTCTCTCTCTC | 299 | 1 | 1.000 | 0.000 |
| RM224 | 11 | (AAG)8(AG)13 | ATCGATCGATCTTCACGAGG | TGCTATAAAAGGCATTCGGG | 157 | 4 | 2.390 | 1.070 |
| RM286 | 11 | (GA)16 | GGCTTCATCTTTGGCGAC | CCGGATTCACGAGATAAACTC | 110 | 4 | 2.484 | 1.066 |
| RM17 | 12 | (GA)21 | TGCCCTGTTATTTTCTTCTCTC | GGTGATCCTTTCCCATTTCA | 184 | 3 | 1.349 | 0.495 |
| RM101 | 12 | (CT)37 | GTGAATGGTCAAGTGACTTAGGTGGC | ACACAACATGTTCCCTCCCATGC | 324 | 3 | 2.563 | 1.001 |
| RM277 | 12 | (GA)11 | CGGTCAAATCATCACCTGAC | CAAGGCTTGCAAGGGAAG | 124 | 2 | 1.929 | 0.675 |
| RM1103 | 12 | (AG)12 | CAGCTGCTGCTACTACACCG | CTACTCCACGTCCATGCATG | 216 | 2 | 1.792 | 0.634 |
| Chr, Chromosome; EPS, Expected PCR product size; Na, Observed number of alleles; Ne, Effective number of alleles; I, Shannon’s information index. | ||||||||
| Origin | Na | Ne | I | No | P (%) | Ho | He | Fis | Fit | Fst | t | |
| Qingpu (Qp), Shanghai, China | 1.833 | 1.560 | 0.422 | 30 | 62.50 | 0.034 | 0.295 | 0.855 | 0.883 | 0.192 | 0.062 | |
| Jiading (Jd), Shanghai, China | 1.458 | 1.300 | 0.228 | 15 | 31.25 | 0.000 | 0.155 | 1.000 | 1.000 | 0.230 | 0.000 | |
| Nanhui (Nh), Shanghai, China | 1.458 | 1.309 | 0.261 | 21 | 43.75 | 0.010 | 0.202 | 0.871 | 0.940 | 0.538 | 0.031 | |
| Pudong (Pd), Shanghai, China | 1.667 | 1.434 | 0.367 | 29 | 60.42 | 0.004 | 0.272 | 0.951 | 0.978 | 0.554 | 0.011 | |
| Chongming (Cm), Shanghai, China | 1.458 | 1.313 | 0.259 | 20 | 41.67 | 0.013 | 0.193 | 0.911 | 0.934 | 0.257 | 0.034 | |
| Fengxian (Fx), Shanghai, China | 1.708 | 1.416 | 0.365 | 29 | 60.42 | 0.050 | 0.262 | 0.708 | 0.785 | 0.263 | 0.120 | |
| Taizhou (Tz), Jiangsu, China | 1.958 | 1.685 | 0.489 | 31 | 64.58 | 0.067 | 0.337 | 0.728 | 0.769 | 0.149 | 0.131 | |
| Shenyang (Sy), Liaoning, China | 1.375 | 1.260 | 0.213 | 17 | 35.42 | 0.008 | 0.158 | 0.893 | 0.939 | 0.424 | 0.031 | |
| Tieling (Tl), Liaoning, China | 1.500 | 1.334 | 0.270 | 20 | 41.67 | 0.004 | 0.195 | 0.963 | 0.976 | 0.340 | 0.012 | |
| Haicheng (Hc), Liaoning, China | 1.500 | 1.308 | 0.266 | 21 | 43.75 | 0.004 | 0.193 | 0.972 | 0.980 | 0.282 | 0.010 | |
| Dandong (Dd), Liaoning, China | 1.458 | 1.349 | 0.269 | 20 | 41.67 | 0.004 | 0.202 | 0.975 | 0.981 | 0.235 | 0.010 | |
| Gongzhuling (Gz), Jilin, China | 1.292 | 1.218 | 0.168 | 12 | 25.00 | 0.000 | 0.124 | 1.000 | 1.000 | 0.241 | 0.000 | |
| Jiamusi (Jm), Heilongjiang, China | 1.438 | 1.337 | 0.258 | 19 | 38.58 | 0.000 | 0.193 | 1.000 | 1.000 | 0.468 | 0.000 | |
| Yesan (Ys), Chungcheong, Korea | 1.458 | 1.269 | 0.231 | 19 | 38.58 | 0.009 | 0.163 | 0.907 | 0.936 | 0.313 | 0.033 | |
| Seosan (Ss), Chungcheong, Korea | 1.542 | 1.27 | 0.267 | 25 | 52.08 | 0.007 | 0.186 | 0.947 | 0.959 | 0.234 | 0.021 | |
| Overall | 3.104 | 2.047 | 0.748 | 43 | 85.58 | 0.012 | 0.434 | 0.931 | 0.973 | 0.606 | 0.014 | |
| Na, Observed number of alleles; Ne, Effective number of alleles; I, Shannon’s diversity index; No, Number of polymorphic loci; P, Percentage of polymorphic loci; Ho, Observed heterozygosity; He, Expected heterozygosity; Fis, Fit and Fst, F-statistics of regional populations referred to | ||||||||||||
Table 4 Genetic characteristics of 76 weedy rice accessions in Northeast Asia by POPGENE according to 48 SSR markers.
| Origin | Na | Ne | I | No | P (%) | Ho | He | Fis | Fit | Fst | t | |
| Qingpu (Qp), Shanghai, China | 1.833 | 1.560 | 0.422 | 30 | 62.50 | 0.034 | 0.295 | 0.855 | 0.883 | 0.192 | 0.062 | |
| Jiading (Jd), Shanghai, China | 1.458 | 1.300 | 0.228 | 15 | 31.25 | 0.000 | 0.155 | 1.000 | 1.000 | 0.230 | 0.000 | |
| Nanhui (Nh), Shanghai, China | 1.458 | 1.309 | 0.261 | 21 | 43.75 | 0.010 | 0.202 | 0.871 | 0.940 | 0.538 | 0.031 | |
| Pudong (Pd), Shanghai, China | 1.667 | 1.434 | 0.367 | 29 | 60.42 | 0.004 | 0.272 | 0.951 | 0.978 | 0.554 | 0.011 | |
| Chongming (Cm), Shanghai, China | 1.458 | 1.313 | 0.259 | 20 | 41.67 | 0.013 | 0.193 | 0.911 | 0.934 | 0.257 | 0.034 | |
| Fengxian (Fx), Shanghai, China | 1.708 | 1.416 | 0.365 | 29 | 60.42 | 0.050 | 0.262 | 0.708 | 0.785 | 0.263 | 0.120 | |
| Taizhou (Tz), Jiangsu, China | 1.958 | 1.685 | 0.489 | 31 | 64.58 | 0.067 | 0.337 | 0.728 | 0.769 | 0.149 | 0.131 | |
| Shenyang (Sy), Liaoning, China | 1.375 | 1.260 | 0.213 | 17 | 35.42 | 0.008 | 0.158 | 0.893 | 0.939 | 0.424 | 0.031 | |
| Tieling (Tl), Liaoning, China | 1.500 | 1.334 | 0.270 | 20 | 41.67 | 0.004 | 0.195 | 0.963 | 0.976 | 0.340 | 0.012 | |
| Haicheng (Hc), Liaoning, China | 1.500 | 1.308 | 0.266 | 21 | 43.75 | 0.004 | 0.193 | 0.972 | 0.980 | 0.282 | 0.010 | |
| Dandong (Dd), Liaoning, China | 1.458 | 1.349 | 0.269 | 20 | 41.67 | 0.004 | 0.202 | 0.975 | 0.981 | 0.235 | 0.010 | |
| Gongzhuling (Gz), Jilin, China | 1.292 | 1.218 | 0.168 | 12 | 25.00 | 0.000 | 0.124 | 1.000 | 1.000 | 0.241 | 0.000 | |
| Jiamusi (Jm), Heilongjiang, China | 1.438 | 1.337 | 0.258 | 19 | 38.58 | 0.000 | 0.193 | 1.000 | 1.000 | 0.468 | 0.000 | |
| Yesan (Ys), Chungcheong, Korea | 1.458 | 1.269 | 0.231 | 19 | 38.58 | 0.009 | 0.163 | 0.907 | 0.936 | 0.313 | 0.033 | |
| Seosan (Ss), Chungcheong, Korea | 1.542 | 1.27 | 0.267 | 25 | 52.08 | 0.007 | 0.186 | 0.947 | 0.959 | 0.234 | 0.021 | |
| Overall | 3.104 | 2.047 | 0.748 | 43 | 85.58 | 0.012 | 0.434 | 0.931 | 0.973 | 0.606 | 0.014 | |
| Na, Observed number of alleles; Ne, Effective number of alleles; I, Shannon’s diversity index; No, Number of polymorphic loci; P, Percentage of polymorphic loci; Ho, Observed heterozygosity; He, Expected heterozygosity; Fis, Fit and Fst, F-statistics of regional populations referred to | ||||||||||||
| Origin | Qp | Jd | Nh | Pd | Cm | Fx | Tz | Sy | Tl | Hc | Dd | Gz | Jm | Ys |
| Jd | 0.078 | |||||||||||||
| Nh | 0.130 | 0.115 | ||||||||||||
| Pd | 0.093 | 0.145 | 0.112 | |||||||||||
| Cm | 0.076 | 0.143 | 0.173 | 0.174 | ||||||||||
| Fx | 0.446 | 0.579 | 0.543 | 0.410 | 0.527 | |||||||||
| Tz | 0.443 | 0.578 | 0.552 | 0.416 | 0.476 | 0.184 | ||||||||
| Sy | 0.738 | 0.889 | 0.832 | 0.622 | 0.777 | 0.254 | 0.145 | |||||||
| Tl | 0.715 | 0.850 | 0.902 | 0.652 | 0.689 | 0.266 | 0.228 | 0.085 | ||||||
| Hc | 0.698 | 0.854 | 0.941 | 0.689 | 0.721 | 0.211 | 0.162 | 0.114 | 0.068 | |||||
| Dd | 0.697 | 0.822 | 0.934 | 0.744 | 0.743 | 0.258 | 0.185 | 0.170 | 0.121 | 0.060 | ||||
| Gz | 0.747 | 0.859 | 0.930 | 0.737 | 0.821 | 0.311 | 0.199 | 0.202 | 0.201 | 0.103 | 0.087 | |||
| Jm | 0.665 | 0.771 | 0.904 | 0.668 | 0.760 | 0.246 | 0.220 | 0.188 | 0.208 | 0.115 | 0.107 | 0.098 | ||
| Ys | 0.721 | 0.849 | 0.863 | 0.687 | 0.828 | 0.228 | 0.220 | 0.157 | 0.180 | 0.115 | 0.138 | 0.106 | 0.053 | |
| Ss | 0.254 | 0.273 | 0.246 | 0.309 | 0.361 | 0.469 | 0.518 | 0.767 | 0.809 | 0.771 | 0.776 | 0.666 | 0.660 | 0.627 |
| Qp, Qingpu, Shanghai, China; Jd, Jiading, Shanghai, China; Nh, Nanhui, Shanghai, China; Pd, Pudong, Shanghai, China; Cm, Chongming, Shanghai, China; Fx, Fengxian, Shanghai, China; Tz, Taizhou, Jiangsu, China; Sy, Shenyang, Liaoning, China; Tl, Tieling, Liaoning, China; Hc, Haicheng, Liaoning, China; Dd, Dandong, Liaoning, China; Gz, Gongzhuling, Jilin, China; Jm, Jiamusi, Heilongjiang, China; Ys, Yesan, Chungcheong, Korea; Ss, Seosan, Chungcheong, Korea. | ||||||||||||||
Table 5 Genetic distance (Nei, 1978) among weedy rice groups in Northeast Asia.
| Origin | Qp | Jd | Nh | Pd | Cm | Fx | Tz | Sy | Tl | Hc | Dd | Gz | Jm | Ys |
| Jd | 0.078 | |||||||||||||
| Nh | 0.130 | 0.115 | ||||||||||||
| Pd | 0.093 | 0.145 | 0.112 | |||||||||||
| Cm | 0.076 | 0.143 | 0.173 | 0.174 | ||||||||||
| Fx | 0.446 | 0.579 | 0.543 | 0.410 | 0.527 | |||||||||
| Tz | 0.443 | 0.578 | 0.552 | 0.416 | 0.476 | 0.184 | ||||||||
| Sy | 0.738 | 0.889 | 0.832 | 0.622 | 0.777 | 0.254 | 0.145 | |||||||
| Tl | 0.715 | 0.850 | 0.902 | 0.652 | 0.689 | 0.266 | 0.228 | 0.085 | ||||||
| Hc | 0.698 | 0.854 | 0.941 | 0.689 | 0.721 | 0.211 | 0.162 | 0.114 | 0.068 | |||||
| Dd | 0.697 | 0.822 | 0.934 | 0.744 | 0.743 | 0.258 | 0.185 | 0.170 | 0.121 | 0.060 | ||||
| Gz | 0.747 | 0.859 | 0.930 | 0.737 | 0.821 | 0.311 | 0.199 | 0.202 | 0.201 | 0.103 | 0.087 | |||
| Jm | 0.665 | 0.771 | 0.904 | 0.668 | 0.760 | 0.246 | 0.220 | 0.188 | 0.208 | 0.115 | 0.107 | 0.098 | ||
| Ys | 0.721 | 0.849 | 0.863 | 0.687 | 0.828 | 0.228 | 0.220 | 0.157 | 0.180 | 0.115 | 0.138 | 0.106 | 0.053 | |
| Ss | 0.254 | 0.273 | 0.246 | 0.309 | 0.361 | 0.469 | 0.518 | 0.767 | 0.809 | 0.771 | 0.776 | 0.666 | 0.660 | 0.627 |
| Qp, Qingpu, Shanghai, China; Jd, Jiading, Shanghai, China; Nh, Nanhui, Shanghai, China; Pd, Pudong, Shanghai, China; Cm, Chongming, Shanghai, China; Fx, Fengxian, Shanghai, China; Tz, Taizhou, Jiangsu, China; Sy, Shenyang, Liaoning, China; Tl, Tieling, Liaoning, China; Hc, Haicheng, Liaoning, China; Dd, Dandong, Liaoning, China; Gz, Gongzhuling, Jilin, China; Jm, Jiamusi, Heilongjiang, China; Ys, Yesan, Chungcheong, Korea; Ss, Seosan, Chungcheong, Korea. | ||||||||||||||
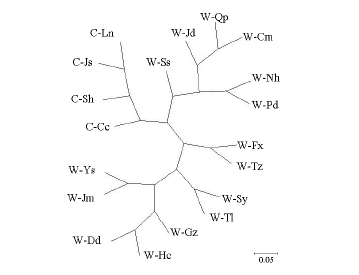
Fig. 2. Unrooted neighbor-joining tree of 15 weedy rice groups and 4 cultivated rice groups by POPGENE, based on a matrix of the inferred genetic diversity. W, Weedy rice; C, Cultivar rice; Qp, Qingpu, Shanghai, China; Jd, Jiading, Shanghai, China; Nh, Nanhui, Shanghai, China; Pd, Pudong, Shanghai, China; Cm, Chongming, Shanghai, China; Fx, Fengxian, Shanghai, China; Tz, Taizhou, Jiangsu, China; Sy, Shenyang, Liaoning, China; Tl, Tieling, Liaoning, China; Hc, Haicheng, Liaoning, China; Dd, Dandong, Liaoning, China; Gz, Gongzhuling, Jilin, China; Jm, Jiamusi, Heilongjiang, China; Ys, Yesan, Chungcheong, Korea; Ss, Seosan, Chungcheong, Korea.
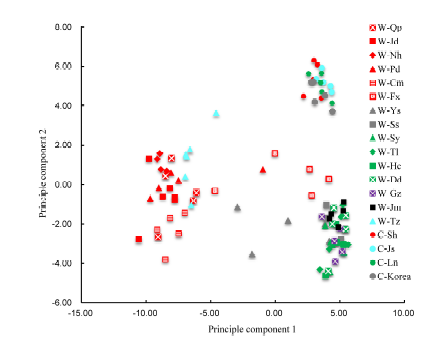
Fig. 4. Two-dimention principle analysis in 96 accessions for Northeast Asia. W, Weedy rice; C, Cultivar rice; Qp, Qingpu, Shanghai, China; Jd, Jiading, Shanghai, China; Nh, Nanhui, Shanghai, China; Pd, Pudong, Shanghai, China; Cm, Chongming, Shanghai, China; Fx, Fengxian, Shanghai, China; Tz, Taizhou, Jiangsu, China; Sy, Shenyang, Liaoning, China; Tl, Tieling, Liaoning, China; Hc, Haicheng, Liaoning, China; Dd, Dandong, Liaoning, China; Gz, Gongzhuling, Jilin, China; Jm, Jiamusi, Heilongjiang, China; Ys, Yesan, Chungcheong, Korea; Ss, Seosan, Chungcheong, Korea.
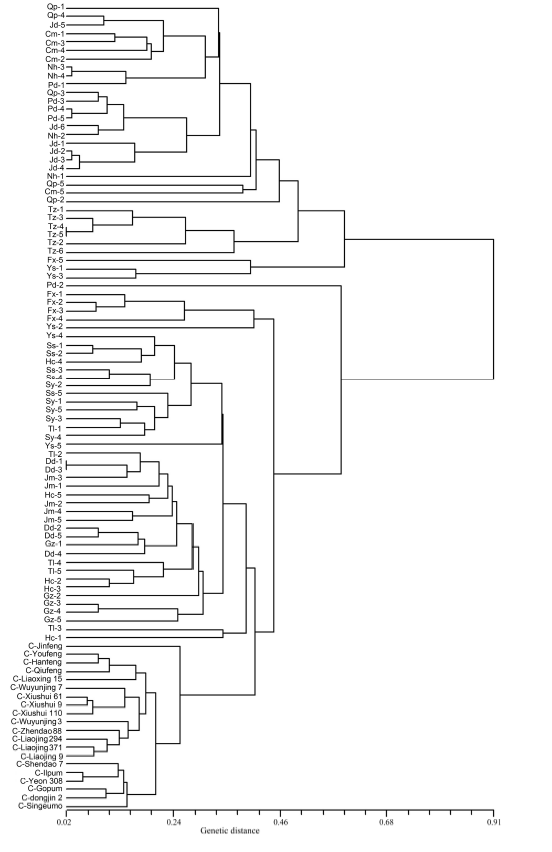

Fig. 3. Cluster diagram of 76 weedy rice and 20 rice accessions in Northeast Asia based on the genetic distance derived from 48 SSR markers, using Nei’s unbiased genetic distance coefficients (Nei, 1978). C, Cultivar rice; Qp, Qingpu, Shanghai, China; Jd, Jiading, Shanghai, China; Nh, Nanhui, Shanghai, China; Pd, Pudong, Shanghai, China; Cm, Chongming, Shanghai, China; Fx, Fengxian, Shanghai, China; Tz, Taizhou, Jiangsu, China; Sy, Shenyang, Liaoning, China; Tl, Tieling, Liaoning, China; Hc, Haicheng, Liaoning, China; Dd, Dandong, Liaoning, China; Gz, Gongzhuling, Jilin, China; Jm, Jiamusi, Heilongjiang, China; Ys, Yesan, Chungcheong, Korea; Ss, Seosan, Chungcheong, Korea.
| 1 | Baker H G.1974. The evolution of weeds.Annu Rev Ecol Syst, 5: 1-24. |
| 2 | Bres-Patry C, Lorieux M, Clément G, Bangratz M, Ghesquière A.2001. Heredity and genetic mapping of domestication-related traits in a temperate japonica weedy rice.Theor Appl Genet, 102(1): 118-126. |
| 3 | Burger J C, Lee S, Ellstrand N C.2006. Origin and genetic structure of feral rye in the western United States.Mol Ecol, 15(9): 2527-2539. |
| 4 | Cao Q J, Lu B R, Xia H, Rong J, Sala F, Spada A, Grassi F.2006. Genetic diversity and origin of weedy rice (Oryza sativa f. spontanea) populations found in north-eastern China revealed by simple sequence repeat (SSR) markers.Ann Bot, 98(6): 1241-1252. |
| 5 | Chen H Z, Xuan S N, Wang W X, Shao G S, Sun Z X.2004. Freezing tolerance and germination ability at low temperature of Dandong weedy rice.Chin J Rice Sci, 18(2): 109-112. (in Chinese with English abstract) |
| 6 | Cho Y C, Chung T Y, Suh H S.1995. Genetic characteristics of Korean weedy rice (Oryza sativa L.) by RFLP analysis.Euphytica, 86(2): 103-110. |
| 7 | Dekker J.1997. Weed diversity and weed management.Weed Sci, 45: 357-363. |
| 8 | Doyle J J, Doyle J L.1987. A rapid DNA isolation procedure for small quantities of flesh leaf tissue.Pytochem Bull, 19: 11-15. |
| 9 | Federici M T, Vaughan D, Tomooka N, Kaga A, Wang X W, Doi K, Francis M, Zorrilla G, Saldain N.2001. Analysis of Uruguayan weedy rice genetic diversity using AFLP molecular markers.Electron J Biotechnol, 4(3): 130-145. |
| 10 | Hartl D L, Clark A G.1989. Principles of Population Genetics. Sunderland, Massachusetts, USA: Sinauer Associate: 682. |
| 11 | Hossain M, Narciso J H.. |
| 12 | Ishikawa R, Toki N, Imai K, Sato Y I, Yamagishi H, Shimamoto Y, Ueno K, Morishima H, Sato T.2005. Origin of weedy rice grown in Bhutan and the force of genetic diversity.Genet Res Crop Evol, 52(4): 395-403. |
| 13 | Jiang H, Wu J L, Wang G L, Wang S.1985. Study on weedy rice from Lianyungang, China.Crop Res China, (2): 4-7. (in Chinese) |
| 14 | Li M B, Ma D R, Xu Z J, Chen W F.2006. Preliminary study on biological trait of Liaoning weedy rice.J Anhui Agric Sci, 34(20): 5224-5225. (in Chinese with English abstract) |
| 15 | Li M B, Ma D R, Xu Z J, Huang C, Piao Z Z, Lee J R, Chen W F.2008. Subspecies characteristics and their relationships with economic characters of weedy rice in Liaoning Province.Liaoning Agric Sci, (4): 1-5. (in Chinese with English abstract) |
| 16 | Li X Y, Qiang S, Song X L, Cai K, Dai W M.2014. Haplotype analysis of Rc gene for weedy rice in Jiangsu Province.Chin J Rice Sci, 28(3): 304-313. (in Chinese with English abstract) |
| 17 | Londo J P, Chiang Y C, Hung K H, Chiang T Y, Schaal B A.2006. Phylogeography of Asian wild rice, Oryza rufipogon, reveals multiple independent domestications of cultivated rice, Oryza sativa.Proc Natl Acad Sci USA, 103(25): 9578-9583. |
| 18 | Ma D R, Chen W F, Xu Z J, Zhang W Z.2005. Origin and control management of weedy rice in Liaoning.Chin Agric Sci Bull, 21(8): 358-360. (in Chinese with English abstract) |
| 19 | Ma D R, Li M B, Wang N, Xu Z J, Chen W F.2008. Genetic diversity and population differentiation of weedy rice in Liaoning Province of China.Acta Agron Sin, 34(3): 403-4l1. (in Chinese with English abstract) |
| 20 | Ma D R, Kong D X, Gao Q, Xu F, Zhao M H, Tang L, Xu Z J, Chen W F.2014. Effects of weedy rice competition on yield and paddy environment of transplanted rice in Northeast China.Chin J Rice Sci, 28(2): 211-216. (in Chinese with English abstract) |
| 21 | Nei M.1978. Estimation of average heterozygosity and genetic distance from a small number of individuals.Genetics, 89(3): 583-590. |
| 22 | Oka H I.1988. Origin of Cultivated Rice. Tokyo: Japan Science Society Press. |
| 23 | Panaud O, Chen X, McCouch S R.1996. Development of microsatellite markers and characterization of simple sequence length polymorphism (SSLP) in rice (Oryza sativa L.).Mol Gen Genet, 252(5): 597-607. |
| 24 | Pyšek P, Prach K.2003. Research into plant invasions in a cross-roads region: History and focus.Biol Invas, 5(4): 337-348. |
| 25 | Rohlf F J.1998. NTSYSpc: Numerical Taxonomy and Multivariate Analysis System, Version 2_02. Setauket, New York: Exeter Software. |
| 26 | Suh H S, Sato Y I, Morishima H.1997. Genetic characterization of weedy rice (Oryza sativa L.) based on morpho-physiology, isozymes and RAPD markers.Theor Appl Genet, 94(3/4): 316-321. |
| 27 | Wright S.1978. Evolution and the Genetics of Population: Variability Within and Among Natural Populations. Chicago: University of Chicago Press. |
| 28 | Xu C, Wu W C.1996. Ecological investigating and identifying of weedy rices in Hainan Island.Chin J Rice Sci, 10(4): 247-249. (in Chinese with English abstract) |
| 29 | Yang S R, Zhao J S.1959. A research on indica-japonica hybridization.Agric Sci, 10: 256-268. (in Chinese) |
| 30 | Yeh F C, Yang R C, Boyle T.1999. Microsoft Window-Based Freeware for Population Genetic Analysis (POPGENE), Version 1.31. |
| 31 | Yu G Q, Bao Y, Shi C H, Dong C Q, Ge S.2005. Genetic diversity and population differentiation of Liaoning weedy rice detected by RAPD and SSR markers.Biochem Genet, 43(5/6): 261-270. |
| 32 | Yu L Q, Mortimer A M, Xuan S N, Lu Y L, Zhou Y J.2005. Stress-resistance of weedy rice Luolijing (Oryza sativa).Chin J Appl Ecol, 16(4): 717-720. (in Chinese with English abstract) |
| 33 | Yuan Q H, Shi L, Wang F, Cao B, Qian Q, Lei X M, Liao Y L, Liu W G, Cheng L, Jia S R.2007. Investigation of rice transgene flow in compass sectors by using male sterile line as a pollen detector.Theor Appl Genet, 115(4): 549-560. |
| 34 | Zhang J, Burgos N R, Ma K, Zhou Y J, Geng R M, Yu L Q.2008. Genetic diversity and relationship of weedy rice in Taizhou City, Jiangsu Province, China.Chin J Rice Sci, 15(4): 295-302. (in Chinese with English abstract) |
| 35 | Zhang Z L, Tan X L, Deng A F.2002. Comprehensive evaluation of weedy rice germplasm.J Plant Genet Res, 3(4): 47-50. (in Chinese with English abstract) |
| 36 | (Managing Editor: Fang Hongmin) |
| [1] | Zhou Ying, Wan Tao, Yuan Bin, Lei Fang, Chen Meijuan, Wang Qiong, Huang Ping, Kou Shuyan, Qiu Wenxiu, Liu Li. Improving Rice Blast Resistance by Mining Broad-Spectrum Resistance Genes at Pik Locus [J]. Rice Science, 2022, 29(2): 133-142. |
| [2] | Guangchen Zhang, Zimeng Liu, Youhong Liu, Noriyuki Kuya, Yuchen Hua, Hongru Shi, Weilin Zhao, Yuqing Han, Toshio Yamamoto, Wenfu Chen, Jian Sun. iTRAQ-Based Proteomics Investigation of Critical Response Proteins in Embryo and Coleoptile During Rice Anaerobic Germination [J]. Rice Science, 2021, 28(4): 391-401. |
| [3] | Pathaichindachote Wanwarang, Panyawut Natjaree, Sikaewtung Kannika, Patarapuwadol Sujin, Muangprom Amorntip. Genetic Diversity and Allelic Frequency of Selected Thai and Exotic Rice Germplasm Using SSR Markers [J]. Rice Science, 2019, 26(6): 393-403. |
| [4] | Donde Ravindra, Kumar Jitendra, Gouda Gayatri, Kumar Gupta Manoj, Mukherjee Mitadru, Yasin Baksh Sk, Mahadani Pradosh, Kumar Sahoo Khirod, Behera Lambodar, Kumar Dash Sushanta. Assessment of Genetic Diversity of Drought Tolerant and Susceptible Rice Genotypes Using Microsatellite Markers [J]. Rice Science, 2019, 26(4): 239-247. |
| [5] | F. Aala Jr Wilson, B. Gregorio Glenn. Morphological and Molecular Characterization of Novel Salt-Tolerant Rice Germplasms from the Philippines and Bangladesh [J]. Rice Science, 2019, 26(3): 178-188. |
| [6] | Khlaimongkhon Sudthana, Chakhonkaen Sriprapai, Pitngam Keasinee, Ditthab Khanittha, Sangarwut Numphet, Panyawut Natjaree, Wasinanon Thiwawan, Mongkolsiriwatana Chareerat, Chunwongse Julapark, Muangprom Amorntip. Molecular Markers and Candidate Genes for Thermo-Sensitive Genic Male Sterile in Rice [J]. Rice Science, 2019, 26(3): 147-156. |
| [7] | Wenqiang Liu, Xiaowu Pan, Yongchao Li, Yonghong Duan, Jun Min, Sanxiong Liu, Licheng Liu, Xinnian Sheng, Xiaoxiang Li. Identification of QTLs and Validation of qCd-2 Associated with Grain Cadmium Concentrations in Rice [J]. Rice Science, 2019, 26(1): 42-49. |
| [8] | Mukherjee Mitadru, Padhy Barada, Srinivasan Bharathkumar, Mahadani Pradosh, Yasin Baksh Sk, Donde Ravindra, Nath Singh Onkar, Behera Lambodar, Swain Padmini, Kumar Dash Sushanta. Revealing Genetic Relationship and Prospecting of Novel Donors Among Upland Rice Genotypes Using qDTY-Linked SSR Markers [J]. Rice Science, 2018, 25(6): 308-319. |
| [9] | Kazemi Sheidollah, Reza Eshghizadeh Hamid, Zahedi Morteza. Responses of Four Rice Varieties to Elevated CO2 and Different Salinity Levels [J]. Rice Science, 2018, 25(3): 142-151. |
| [10] | Radhesh Krishnan Subramanian, Muthuramalingam Pandiyan, Pandian Subramani, Banupriya Ramachandradoss, Chithra Gunasekar, Ramesh Manikandan. Sprouted Sorghum Extract Elicits Coleoptile Emergence, Enhances Shoot and Root Acclimatization, and Maintains Genetic Fidelity in indica Rice [J]. Rice Science, 2018, 25(2): 61-72. |
| [11] | Xiongsiyee Vua, Rerkasem Benjavan, Veeradittakit Jeeraporn, Saenchai Chorpet, Lordkaew Sittichai, Thebault Prom-u-thai Chanakan. Variation in Grain Quality of Upland Rice from Luang Prabang Province, Lao PDR [J]. Rice Science, 2018, 25(2): 94-102. |
| [12] | Haritha G., P. M. Swamy B., L. Naik M., Jyothi B., Divya B., Malathi S., Sarla N.. Yield Traits and Associated Marker Segregation in Elite Introgression Lines Derived from O. sativa × O. nivara [J]. Rice Science, 2018, 25(1): 19-31. |
| [13] | Naga Bheema Lingeswara Reddy Inja, Kim Sung-Mi, Kim Beom-Ki, Yoon In-Sun, Kwon Taek-Ryoun. Identification of Rice Accessions Associated with K+/Na+ Ratio and Salt Tolerance Based on Physiological and Molecular Responses [J]. Rice Science, 2017, 24(6): 360-364. |
| [14] | Rekha Talukdar Preeti, Rathi Sunayana, Pathak Khanin, Kumar Chetia Sanjay, Nath Sarma Ramendra. Population Structure and Marker-Trait Association in Indigenous Aromatic Rice [J]. Rice Science, 2017, 24(3): 145-154. |
| [15] | Anupam Alpana, Imam Jahangir, Mohammad Quatadah Syed, Siddaiah Anantha, Prasad Das Shankar, Variar Mukund, Prasad Mandal Nimai. Genetic Diversity Analysis of Rice Germplasm in Tripura State of Northeast India Using Drought and Blast Linked Markers [J]. Rice Science, 2017, 24(1): 10-20. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||