
Rice Science ›› 2016, Vol. 23 ›› Issue (4): 211-218.DOI: 10.1016/j.rsci.2016.06.004
• Orginal Article • Previous Articles Next Articles
Sharf Uddin Ahmed Mir1, Khalequzzaman Mohammad1, Khairul Bashar Md.2, Kalam Mohammad Shamsuddin Abul3
Received:2015-06-08
Accepted:2015-12-14
Online:2016-07-28
Published:2016-04-11
Sharf Uddin Ahmed Mir, Khalequzzaman Mohammad, Khairul Bashar Md., Kalam Mohammad Shamsuddin Abul. Agro-Morphological, Physico-Chemical and Molecular Characterization of Rice Germplasm with Similar Names of Bangladesh[J]. Rice Science, 2016, 23(4): 211-218.
Add to citation manager EndNote|Ris|BibTeX
| Serial No. | Accession name | Accession No. a | Origin |
|---|---|---|---|
| KS1 | Kartik Sail | 3243 | Sylhet |
| KS2 | Kartik Sail | 776 | Chittagong |
| KS3 | Katih Shail | 438 | Rajshahi |
| KS4 | Kartik Sail | 539 | Rangpur |
| KS5 | Kartik Sail | 77 | Dhaka |
| KS6 | Kati Shail | 170 | Tangail |
| KS7 | Kartik Sail | 3662 | Sherpur |
| KS8 | Kati Shail | 3631 | Rajshahi |
| KS9 | Kartika | 4053 | Sylhet |
| KS10 | Kartik Sail | 4881 | Tangail |
| KS11 | Kartik Sail | 76 | Dhaka |
| KS12 | Kati Shail | 437 | Rajshahi |
| KS13 | Kartik Sail | 78 | Dhaka |
| KS14 | Kartik Sail | 1882 | Kishorganj |
| KS15 | Kartik sail (2) | 689 | Comilla |
| KS16 | Kartik sail (2) | 846 | Sylhet |
| KS17 | Kartik Sail | 664 | Comilla |
| KS18 | Kartik Sail | 1887 | Kishorganj |
| KS19 | Kartik Sail | 844 | Sylhet |
| KS20 | Katih Shail | 994 | Khulna |
| KS21 | Kartik Sail | 845 | Sylhet |
| DB1 | Dhali Boro | 2250 | Sylhet |
| DB2 | Dhali Boro | 2247 | Sylhet |
| DB3 | Dhali Boro | 2249 | Sylhet |
| DB4 | Dholi Boro | 180 | Tangail |
| DB5 | Dhali Boro | 2245 | Sylhet |
| DB6 | Dholi Boro | 4396 | Sylhet |
| DB7 | Dhali Boro | 2246 | Sylhet |
| DB8 | Dhali Boro | 2244 | Sylhet |
| DB9 | Dhali Boro | 2248 | Sylhet |
| DB10 | Dhali Boro | 2243 | Sylhet |
Table 1 List of 31 duplicate and similar named rice germplasms of Bangladesh.
| Serial No. | Accession name | Accession No. a | Origin |
|---|---|---|---|
| KS1 | Kartik Sail | 3243 | Sylhet |
| KS2 | Kartik Sail | 776 | Chittagong |
| KS3 | Katih Shail | 438 | Rajshahi |
| KS4 | Kartik Sail | 539 | Rangpur |
| KS5 | Kartik Sail | 77 | Dhaka |
| KS6 | Kati Shail | 170 | Tangail |
| KS7 | Kartik Sail | 3662 | Sherpur |
| KS8 | Kati Shail | 3631 | Rajshahi |
| KS9 | Kartika | 4053 | Sylhet |
| KS10 | Kartik Sail | 4881 | Tangail |
| KS11 | Kartik Sail | 76 | Dhaka |
| KS12 | Kati Shail | 437 | Rajshahi |
| KS13 | Kartik Sail | 78 | Dhaka |
| KS14 | Kartik Sail | 1882 | Kishorganj |
| KS15 | Kartik sail (2) | 689 | Comilla |
| KS16 | Kartik sail (2) | 846 | Sylhet |
| KS17 | Kartik Sail | 664 | Comilla |
| KS18 | Kartik Sail | 1887 | Kishorganj |
| KS19 | Kartik Sail | 844 | Sylhet |
| KS20 | Katih Shail | 994 | Khulna |
| KS21 | Kartik Sail | 845 | Sylhet |
| DB1 | Dhali Boro | 2250 | Sylhet |
| DB2 | Dhali Boro | 2247 | Sylhet |
| DB3 | Dhali Boro | 2249 | Sylhet |
| DB4 | Dholi Boro | 180 | Tangail |
| DB5 | Dhali Boro | 2245 | Sylhet |
| DB6 | Dholi Boro | 4396 | Sylhet |
| DB7 | Dhali Boro | 2246 | Sylhet |
| DB8 | Dhali Boro | 2244 | Sylhet |
| DB9 | Dhali Boro | 2248 | Sylhet |
| DB10 | Dhali Boro | 2243 | Sylhet |
| Cluster | No. of genotypes | Serial No. |
|---|---|---|
| I | 11 | KS1, KS2, KS5, KS7, KS9, KS11, KS14, KS17, KS18, KS19, KS21 |
| II | 3 | KS6, KS8, KS13 |
| III | 7 | KS3, KS4, KS10, KS12, KS15, KS16, KS20 |
| IV | 10 | DB1, DB2, DB3, DB4, DB5, DB6, DB7, DB8, DB9, DB10 |
Table 2 Distribution of 31 duplicate and similar named rice germplasms into 4 clusters for 18 morpho-physicochemical characters.
| Cluster | No. of genotypes | Serial No. |
|---|---|---|
| I | 11 | KS1, KS2, KS5, KS7, KS9, KS11, KS14, KS17, KS18, KS19, KS21 |
| II | 3 | KS6, KS8, KS13 |
| III | 7 | KS3, KS4, KS10, KS12, KS15, KS16, KS20 |
| IV | 10 | DB1, DB2, DB3, DB4, DB5, DB6, DB7, DB8, DB9, DB10 |
| Cluster | I | II | III | IV |
|---|---|---|---|---|
| I | 0.93 | |||
| II | 6.91 | 0.69 | ||
| III | 5.07 | 10.21 | 0.74 | |
| IV | 24.61 | 22.17 | 22.06 | 0.62 |
Table 3 Average intra- (bold) and inter-cluster distances (D2) for 18 morpho-physicochemical characters of 31 duplicate and similar named rice germplasms.
| Cluster | I | II | III | IV |
|---|---|---|---|---|
| I | 0.93 | |||
| II | 6.91 | 0.69 | ||
| III | 5.07 | 10.21 | 0.74 | |
| IV | 24.61 | 22.17 | 22.06 | 0.62 |
| Character | Cluster I | Cluster II | Cluster III | Cluster IV | Vector I | Vector II | Combined ranking a |
|---|---|---|---|---|---|---|---|
| Seedling height (cm) | 66 | 69.9 | 56.7 | 34.8 | 0.1093 | -0.0729 | 9 |
| Plant height (cm) | 123.9 | 125.6 | 111.1 | 106.1 | -0.1082 | 0.0332 | 12 |
| Culm diameter (mm) | 5.27 | 5.16 | 4.77 | 4.28 | 1.7199 | -1.788 | 11 |
| Panicle exsertion (mm) | 0.64 | 0.53 | 0.44 | 6.13 | -1.6266 | -0.2472 | 18 |
| Effective tillering number per hill | 11.8 | 11 | 12.1 | 10.3 | 0.2065 | 0.2738 | 3 |
| Days to maturity (d) | 143 | 141 | 142 | 160 | -0.2783 | -0.1692 | 14 |
| Primary branch number per panicle | 10.6 | 9 | 9.1 | 7.1 | 0.847 | 0.1551 | 2 |
| Secondary branch number per panicle | 33.6 | 21.7 | 24.9 | 16.9 | 0.1115 | 0.24 | 6 |
| Grain yield per panicle (g) | 3.2 | 2.7 | 2.3 | 1.9 | 0.2634 | -1.1171 | 17 |
| Grain length (mm) | 8.55 | 8.44 | 8.17 | 7.62 | 0.2474 | -0.3449 | 13 |
| Awn length (mm) | 0.6 | 21.5 | 0 | 30.5 | -0.0198 | 0.1587 | 7 |
| 1000-grain weight (g) | 23.19 | 27.18 | 20.82 | 20.69 | -0.0652 | 0.5106 | 4 |
| Grain yield per hill (g) | 22.42 | 21.28 | 22 | 17.28 | -0.3271 | -0.1641 | 15 |
| Biological yield (g) | 40.9 | 37 | 33.1 | 27.7 | 0.1759 | 0.2617 | 5 |
| Milling rate (%) | 68.2 | 65.3 | 69.9 | 68.7 | 0.0361 | -0.633 | 16 |
| Cooking time (min) | 17 | 20 | 16.5 | 19.1 | -0.4281 | 0.3856 | 10 |
| Amylose content (%) | 25.3 | 25.27 | 25.76 | 21.17 | 0.2999 | -0.2623 | 8 |
| Protein content (%) | 7.65 | 9.03 | 7.92 | 9.12 | 0.8121 | 0.6391 | 1 |
Table 4 Means and latent vectors for 18 morpho-physicochemical characters of 31 duplicate and similar named rice germplasms.
| Character | Cluster I | Cluster II | Cluster III | Cluster IV | Vector I | Vector II | Combined ranking a |
|---|---|---|---|---|---|---|---|
| Seedling height (cm) | 66 | 69.9 | 56.7 | 34.8 | 0.1093 | -0.0729 | 9 |
| Plant height (cm) | 123.9 | 125.6 | 111.1 | 106.1 | -0.1082 | 0.0332 | 12 |
| Culm diameter (mm) | 5.27 | 5.16 | 4.77 | 4.28 | 1.7199 | -1.788 | 11 |
| Panicle exsertion (mm) | 0.64 | 0.53 | 0.44 | 6.13 | -1.6266 | -0.2472 | 18 |
| Effective tillering number per hill | 11.8 | 11 | 12.1 | 10.3 | 0.2065 | 0.2738 | 3 |
| Days to maturity (d) | 143 | 141 | 142 | 160 | -0.2783 | -0.1692 | 14 |
| Primary branch number per panicle | 10.6 | 9 | 9.1 | 7.1 | 0.847 | 0.1551 | 2 |
| Secondary branch number per panicle | 33.6 | 21.7 | 24.9 | 16.9 | 0.1115 | 0.24 | 6 |
| Grain yield per panicle (g) | 3.2 | 2.7 | 2.3 | 1.9 | 0.2634 | -1.1171 | 17 |
| Grain length (mm) | 8.55 | 8.44 | 8.17 | 7.62 | 0.2474 | -0.3449 | 13 |
| Awn length (mm) | 0.6 | 21.5 | 0 | 30.5 | -0.0198 | 0.1587 | 7 |
| 1000-grain weight (g) | 23.19 | 27.18 | 20.82 | 20.69 | -0.0652 | 0.5106 | 4 |
| Grain yield per hill (g) | 22.42 | 21.28 | 22 | 17.28 | -0.3271 | -0.1641 | 15 |
| Biological yield (g) | 40.9 | 37 | 33.1 | 27.7 | 0.1759 | 0.2617 | 5 |
| Milling rate (%) | 68.2 | 65.3 | 69.9 | 68.7 | 0.0361 | -0.633 | 16 |
| Cooking time (min) | 17 | 20 | 16.5 | 19.1 | -0.4281 | 0.3856 | 10 |
| Amylose content (%) | 25.3 | 25.27 | 25.76 | 21.17 | 0.2999 | -0.2623 | 8 |
| Protein content (%) | 7.65 | 9.03 | 7.92 | 9.12 | 0.8121 | 0.6391 | 1 |
| Chromosome | Marker | Position | No. of alleles | Size | Major allele | No. of | No. of | Gene | Polymorphism information content | ||
|---|---|---|---|---|---|---|---|---|---|---|---|
| (cM) | (bp) | Size (bp) | Frequency (%) | rare alleles | null alleles | diversity | |||||
| 1 | RM283 | 31.4 | 5 | 141-150 | 150 | 44 | 4 | 1 | 0.62 | 0.55 | |
| RM259 | 54.2 | 6 | 166-187 | 170 | 39 | 4 | 0 | 0.76 | 0.73 | ||
| RM237 | 115.2 | 6 | 116-133 | 130 | 33 | 3 | 1 | 0.78 | 0.75 | ||
| RM302 | 147.8 | 9 | 107-214 | 178 | 31 | 7 | 2 | 0.82 | 0.8 | ||
| RM154 | 4.8 | 8 | 157-183 | 163/167 | 22 | 6 | 0 | 0.84 | 0.82 | ||
| 2 | RM279 | 17.3 | 8 | 135-173 | 166 | 33 | 6 | 3 | 0.82 | 0.8 | |
| RM324 | 66 | 9 | 132-166 | 166 | 28 | 7 | 0 | 0.82 | 0.8 | ||
| RM250 | 170.1 | 5 | 149-155 | 152 | 61 | 4 | 0 | 0.59 | 0.56 | ||
| RM60 | 0 | 10 | 147-182 | 162 | 25 | 8 | 1 | 0.86 | 0.85 | ||
| 3 | RM218 | 67.8 | 10 | 126-152 | 136 | 22 | 8 | 3 | 0.86 | 0.84 | |
| RM55 | 168.2 | 10 | 215-240 | 229 | 17 | 9 | 3 | 0.89 | 0.88 | ||
| RM227 | 214.7 | 4 | 97-105 | 97 | 47 | 1 | 1 | 0.66 | 0.6 | ||
| RM307 | 0 | 6 | 123-177 | 131 | 69 | 5 | 3 | 0.5 | 0.48 | ||
| 4 | RM273 | 94.4 | 7 | 200-217 | 202 | 39 | 4 | 0 | 0.77 | 0.74 | |
| RM241 | 106.2 | 7 | 100-130 | 126 | 44 | 5 | 0 | 0.72 | 0.69 | ||
| RM127 | 150.1 | 4 | 212-224 | 218 | 56 | 1 | 0 | 0.61 | 0.56 | ||
| RM413 | 26.7 | 9 | 68-105 | 68 | 22 | 7 | 2 | 0.86 | 0.85 | ||
| 5 | RM267 | 28.6 | 8 | 150-185 | 166 | 36 | 6 | 0 | 0.75 | 0.71 | |
| RM161 | 96.9 | 6 | 166-182 | 171 | 36 | 3 | 0 | 0.77 | 0.73 | ||
| RM133 | 0 | 4 | 225-232 | 227 | 69 | 2 | 0 | 0.48 | 0.44 | ||
| 6 | RM584 | 26.2 | 10 | 160-214 | 163 | 33 | 8 | 0 | 0.83 | 0.81 | |
| RM541 | 75.5 | 11 | 142-194 | 160 | 25 | 9 | 0 | 0.84 | 0.83 | ||
| RM162 | 108.3 | 6 | 223-233 | 223 | 33 | 3 | 0 | 0.77 | 0.74 | ||
| RM125 | 24.8 | 4 | 130-145 | 141 | 61 | 1 | 0 | 0.56 | 0.51 | ||
| 7 | RM214 | 34.7 | 8 | 110-137 | 117 | 33 | 5 | 0 | 0.78 | 0.75 | |
| RM18 | 90.4 | 8 | 160-180 | 169 | 22 | 5 | 1 | 0.85 | 0.84 | ||
| RM337 | 1.1 | 6 | 162-202 | 168 | 42 | 3 | 0 | 0.71 | 0.67 | ||
| 8 | RM223 | 80.5 | 8 | 138-163 | 155 | 31 | 5 | 0 | 0.82 | 0.8 | |
| RM256 | 101.5 | 8 | 103-138 | 108 | 36 | 5 | 0 | 0.78 | 0.76 | ||
| RM433 | 116 | 13 | 207-248 | 237 | 28 | 12 | 1 | 0.87 | 0.86 | ||
| RM296 | 0 | 8 | 121-139 | 125 | 39 | 6 | 0 | 0.72 | 0.68 | ||
| 9 | RM242 | 73.3 | 8 | 190-214 | 203 | 28 | 6 | 0 | 0.83 | 0.81 | |
| RM215 | 99.4 | 7 | 159-176 | 163 | 31 | 6 | 1 | 0.83 | 0.81 | ||
| RM311 | 25.2 | 10 | 170-201 | 176/182 | 19 | 7 | 0 | 0.86 | 0.84 | ||
| 10 | RM271 | 59.4 | 8 | 81-98 | 93 | 28 | 6 | 1 | 0.82 | 0.8 | |
| RM258 | 70.8 | 13 | 139-167 | 146 | 22 | 11 | 1 | 0.88 | 0.87 | ||
| RM171 | 92.8 | 6 | 314-340 | 314 | 36 | 4 | 0 | 0.75 | 0.72 | ||
| RM21 | 85.7 | 14 | 129-173 | 166 | 22 | 12 | 0 | 0.89 | 0.88 | ||
| 11 | RM229 | 77.8 | 8 | 105-129 | 112 | 25 | 5 | 0 | 0.82 | 0.8 | |
| RM206 | 102.9 | 13 | 116-171 | 157 | 17 | 12 | 0 | 0.9 | 0.89 | ||
| RM224 | 120.1 | 9 | 123-155 | 147 | 36 | 7 | 0 | 0.8 | 0.78 | ||
| RM286 | 0 | 6 | 96-121 | 96 | 31 | 4 | 0 | 0.79 | 0.75 | ||
| 12 | RM19 | 20.9 | 8 | 223-239 | 228 | 28 | 5 | 1 | 0.83 | 0.81 | |
| RM247 | 32.3 | 6 | 126-144 | 136 | 31 | 3 | 0 | 0.78 | 0.75 | ||
| RM277 | 57.2 | 3 | 116-123 | 116/119 | 47 | 1 | 0 | 0.55 | 0.45 | ||
| Minimum | 3 | 68 | 17 | 1 | 0 | 0.48 | 0.44 | ||||
| Average | 7.8 | 161.2 | 35 | 5.8 | 0.6 | 0.77 | 0.74 | ||||
| Maximum | 14 | 314 | 69 | 12 | 3 | 0.9 | 0.89 | ||||
Table 5 Allele variation, gene diversity and polymorphism information content of 45 simple sequence repeat markers across 31 duplicate and similar named rice germplasms.
| Chromosome | Marker | Position | No. of alleles | Size | Major allele | No. of | No. of | Gene | Polymorphism information content | ||
|---|---|---|---|---|---|---|---|---|---|---|---|
| (cM) | (bp) | Size (bp) | Frequency (%) | rare alleles | null alleles | diversity | |||||
| 1 | RM283 | 31.4 | 5 | 141-150 | 150 | 44 | 4 | 1 | 0.62 | 0.55 | |
| RM259 | 54.2 | 6 | 166-187 | 170 | 39 | 4 | 0 | 0.76 | 0.73 | ||
| RM237 | 115.2 | 6 | 116-133 | 130 | 33 | 3 | 1 | 0.78 | 0.75 | ||
| RM302 | 147.8 | 9 | 107-214 | 178 | 31 | 7 | 2 | 0.82 | 0.8 | ||
| RM154 | 4.8 | 8 | 157-183 | 163/167 | 22 | 6 | 0 | 0.84 | 0.82 | ||
| 2 | RM279 | 17.3 | 8 | 135-173 | 166 | 33 | 6 | 3 | 0.82 | 0.8 | |
| RM324 | 66 | 9 | 132-166 | 166 | 28 | 7 | 0 | 0.82 | 0.8 | ||
| RM250 | 170.1 | 5 | 149-155 | 152 | 61 | 4 | 0 | 0.59 | 0.56 | ||
| RM60 | 0 | 10 | 147-182 | 162 | 25 | 8 | 1 | 0.86 | 0.85 | ||
| 3 | RM218 | 67.8 | 10 | 126-152 | 136 | 22 | 8 | 3 | 0.86 | 0.84 | |
| RM55 | 168.2 | 10 | 215-240 | 229 | 17 | 9 | 3 | 0.89 | 0.88 | ||
| RM227 | 214.7 | 4 | 97-105 | 97 | 47 | 1 | 1 | 0.66 | 0.6 | ||
| RM307 | 0 | 6 | 123-177 | 131 | 69 | 5 | 3 | 0.5 | 0.48 | ||
| 4 | RM273 | 94.4 | 7 | 200-217 | 202 | 39 | 4 | 0 | 0.77 | 0.74 | |
| RM241 | 106.2 | 7 | 100-130 | 126 | 44 | 5 | 0 | 0.72 | 0.69 | ||
| RM127 | 150.1 | 4 | 212-224 | 218 | 56 | 1 | 0 | 0.61 | 0.56 | ||
| RM413 | 26.7 | 9 | 68-105 | 68 | 22 | 7 | 2 | 0.86 | 0.85 | ||
| 5 | RM267 | 28.6 | 8 | 150-185 | 166 | 36 | 6 | 0 | 0.75 | 0.71 | |
| RM161 | 96.9 | 6 | 166-182 | 171 | 36 | 3 | 0 | 0.77 | 0.73 | ||
| RM133 | 0 | 4 | 225-232 | 227 | 69 | 2 | 0 | 0.48 | 0.44 | ||
| 6 | RM584 | 26.2 | 10 | 160-214 | 163 | 33 | 8 | 0 | 0.83 | 0.81 | |
| RM541 | 75.5 | 11 | 142-194 | 160 | 25 | 9 | 0 | 0.84 | 0.83 | ||
| RM162 | 108.3 | 6 | 223-233 | 223 | 33 | 3 | 0 | 0.77 | 0.74 | ||
| RM125 | 24.8 | 4 | 130-145 | 141 | 61 | 1 | 0 | 0.56 | 0.51 | ||
| 7 | RM214 | 34.7 | 8 | 110-137 | 117 | 33 | 5 | 0 | 0.78 | 0.75 | |
| RM18 | 90.4 | 8 | 160-180 | 169 | 22 | 5 | 1 | 0.85 | 0.84 | ||
| RM337 | 1.1 | 6 | 162-202 | 168 | 42 | 3 | 0 | 0.71 | 0.67 | ||
| 8 | RM223 | 80.5 | 8 | 138-163 | 155 | 31 | 5 | 0 | 0.82 | 0.8 | |
| RM256 | 101.5 | 8 | 103-138 | 108 | 36 | 5 | 0 | 0.78 | 0.76 | ||
| RM433 | 116 | 13 | 207-248 | 237 | 28 | 12 | 1 | 0.87 | 0.86 | ||
| RM296 | 0 | 8 | 121-139 | 125 | 39 | 6 | 0 | 0.72 | 0.68 | ||
| 9 | RM242 | 73.3 | 8 | 190-214 | 203 | 28 | 6 | 0 | 0.83 | 0.81 | |
| RM215 | 99.4 | 7 | 159-176 | 163 | 31 | 6 | 1 | 0.83 | 0.81 | ||
| RM311 | 25.2 | 10 | 170-201 | 176/182 | 19 | 7 | 0 | 0.86 | 0.84 | ||
| 10 | RM271 | 59.4 | 8 | 81-98 | 93 | 28 | 6 | 1 | 0.82 | 0.8 | |
| RM258 | 70.8 | 13 | 139-167 | 146 | 22 | 11 | 1 | 0.88 | 0.87 | ||
| RM171 | 92.8 | 6 | 314-340 | 314 | 36 | 4 | 0 | 0.75 | 0.72 | ||
| RM21 | 85.7 | 14 | 129-173 | 166 | 22 | 12 | 0 | 0.89 | 0.88 | ||
| 11 | RM229 | 77.8 | 8 | 105-129 | 112 | 25 | 5 | 0 | 0.82 | 0.8 | |
| RM206 | 102.9 | 13 | 116-171 | 157 | 17 | 12 | 0 | 0.9 | 0.89 | ||
| RM224 | 120.1 | 9 | 123-155 | 147 | 36 | 7 | 0 | 0.8 | 0.78 | ||
| RM286 | 0 | 6 | 96-121 | 96 | 31 | 4 | 0 | 0.79 | 0.75 | ||
| 12 | RM19 | 20.9 | 8 | 223-239 | 228 | 28 | 5 | 1 | 0.83 | 0.81 | |
| RM247 | 32.3 | 6 | 126-144 | 136 | 31 | 3 | 0 | 0.78 | 0.75 | ||
| RM277 | 57.2 | 3 | 116-123 | 116/119 | 47 | 1 | 0 | 0.55 | 0.45 | ||
| Minimum | 3 | 68 | 17 | 1 | 0 | 0.48 | 0.44 | ||||
| Average | 7.8 | 161.2 | 35 | 5.8 | 0.6 | 0.77 | 0.74 | ||||
| Maximum | 14 | 314 | 69 | 12 | 3 | 0.9 | 0.89 | ||||
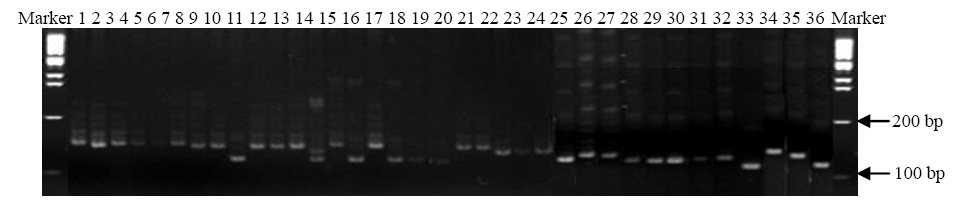
Fig. 2. DNA profile of 36 rice genotypes using RM224 marker.1, KS1; 2, KS2; 3, KS3; 4, KS4; 5, KS5; 6, KS6; 7, KS7; 8, KS8; 9, KS9; 10, KS10; 11, KS11; 12, KS12; 13, KS13; 14, KS14; 15, KS15; 16, KS16; 17, KS17; 18, KS18; 19, KS19; 20, KS20; 21, KS21; 22, BR4; 23, BR23; 24, DB1; 25, DB2; 26, DB3; 27, DB4; 28, DB5; 29, DB6; 30, DB7; 31, DB8; 32, DB9; 33, DB10; 34, BRRI dhan 28; 35, BRRI dhan 29; 36, BR14.
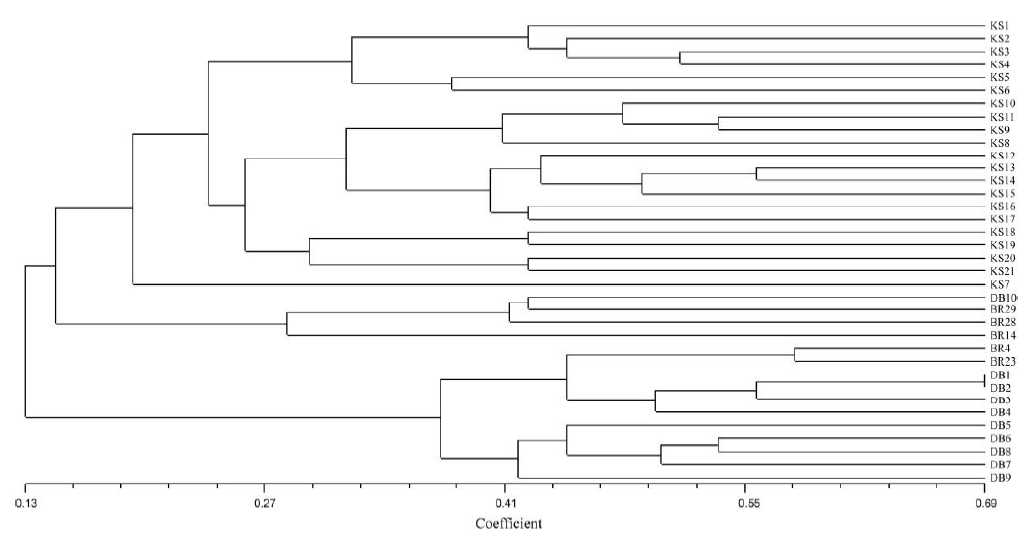
Fig. 3. Dendrogram of 31 duplicate and similar named rice germplasms derived from unweighted pair-group method with arithmetic means cluster analysis based on Nei’s genetic distance across 45 simple sequence repeat markers.BR28, BRRI dhan 28; BR29, BRRI dhan 29.
| 1 | Adair C R.1952. The Mcgill miller method for determining the milling quality of small samples of rice.Rice J, 55(2): 21-23. |
| 2 | Ahmed M S, Akter K, Khalequzzaman M, Rashid E S M H, Bashar M K.2010. Diversity analysis in Boro rice (Oryza sativa L.) accessions.Bangl J Agric Res, 35(1): 29-36. |
| 3 | AOAC.1995. Association of Official Agricultural Chemists. Methods of Analysis. Washington DC, USA. |
| 4 | Chakravorty A, Ghosh P D, Sahu P K.2013. Multivariate analysis of phenotypic diversity of land races of rice of West Bengal.Am J Exp Agric, 3(1): 110-123. |
| 5 | Chen X, Cho Y G, McCouch S R.2002. Sequence divergence of rice microsatellites in Oryza and other plant species.Mol Genet Genom, 268: 331-343. |
| 6 | Collard B C Y, Das A, Virk P S, Mackill D J.2007. Evaluation of quick and dirty DNA extraction methods for marker-assisted selection in rice (Oryza sativa L.).Plant Breeding, 126(1): 47-50. |
| 7 | Datt S, Mani S C.2003. Genetic divergence in elite genotypes of Basmati rice (Oryza sativa L.).Ind J Genet Plant Breeding, 63(1): 73-74. |
| 8 | Fukuoka S, Suu T D, Ebanna K, Trinh L N, Nagamine T, Okuno K.2006. Diversity in phenotypic profiles in land races populations of Vietnamese rice: A case study of agronomic characters for conserving crop genetic diversity on farm.Genet Resour Crop Evol, 53: 753-761. |
| 9 | Hamid A, Nasiruddin M, Haque M, Haque E.1982. Collection of names of the local varieties. In: Taluckdar M H R. Local rice varieties. Joydevpur, Gazipur, Bangladesh: Bangladesh Rice Research Institute. (in Bengali) |
| 10 | Hossain M Z.2008. Genetic diversity study in fine grain and aromatic land races of rice (Oryza sativa L.) by morpho-physico-chemical characters and microsatellite DNA markers. Gazipur, Bangladesh: Bangabandhu Sheikh Mujibur Rahman Agricultual University. |
| 11 | Hossain M, Jaim W M H, Alam M S, Rahman A N M M.2013. Rice biodiversity in Bangladesh: Adoption, diffusion and disappearance of varieties. Dhaka, Bangladesh: BRAC Research and Evalution Division. |
| 12 | Irish B M, Goenaga R, Zhang D, Schnell R J, Brown J S, Motamayor J C.2010. Microsatellite fingerprinting of the USDA-ARS tropical agriculture research station cacao (Theobroma cacao L.) germplasm collection.Crop Sci, 50: 656-667. |
| 13 | Rholf F.2002. NTSYS-pc: Numerical Taxonomy and Multivariate Analysis System Version 2.2. New York, USA: Department of Ecology and Evolution, State University of New York. |
| 14 | Joshi R K, Behera L.2006. Identification and differentiation of indigenous non-Basmati aromatic rice genotypes of India using microsatellite markers.Afr J Biotechnol, 6(4): 348-354. |
| 15 | Juilano B O, Nazareno M B, Ramos N.1969. Properties of waxy and isogenic nonwaxy rices differing in starch gelatinization temperature.J Agric Food Chem, 17: 1364-1369. |
| 16 | Juilano B O.1971. A simplified assay for milled rice amylose.Cereal Sci Today, 16: 334-360. |
| 17 | Junjian N I, Colowit P M, Mackill D J.2002. Evaluation of genetic diversity in rice subspecies using microsatellite markers.Crop Sci, 42: 601-607. |
| 18 | Kaushik A, Jain S, McCouch S R, Jain R K.2011. Phylogenetic relationships among various groups of rice (Oryza sativa L.) as revealed by microsatellite and transposable element-based marker analysis.Ind J Genet Plant Breeding, 71(2): 139-150. |
| 19 | Liu K, Muse S V.2005. Power Marker: Integrated analysis environment for genetic marker data.Bioinformatics, 21: 2128-2129. |
| 20 | Lund B, Ortiz R, Skovgaard I M, Waugh R, Andersen S B.2003. Analysis of potential duplicates in barley genebank collections using re-sampling of microsatellite data.Theor Appl Genet, 106(6): 1129-1138. |
| 21 | Masuduzzaman M.2010. Morphological, molecular characterization and Agrobactrium-mediated genetic transformation in jute (Corchorus spp). Mymensingh, Bangladesh: Bangladesh Agricultural University: 41-46. |
| 22 | Nandakumar N, Singh A K, Sharma R K, Mohapatra T, Prabhu K V, Zaman F U.2004. Molecular fingerprinting of hybrids and assessment of genetic purity of hybrid seeds in rice using microsatellite markers.Euphytica, 136(3): 257-264. |
| 23 | Nascimento W F, Silva E F, Veasey E A.2011. Agro- morphological characterization of upland rice accessions.Sci Agric, 68(6): 652-660. |
| 24 | Nei M, Takezaki N.1994. Estimation of genetic distances and phylogenetic trees from DNA analysis. Proceedings, 5th World Congress on Genetics Applied to Livestock Production, University of Guelph, Guelph, Ontario, Canada, 7-12 August. |
| 25 | Rao C R.1952. Advance Statistical Methods in Biometrical Research. New York, USA: John Willey and Sons Inc: 390. |
| 26 | Rao C R.1964. The use and interception of principal component analysis in applied research.Sankhya Ser A, 22: 317-318. |
| 27 | Roy S K, Kundu A, Chand S P, Senapati B K.2004. Diversity of panicle characters in Aman rice (Oryza sativa L.).Environ Ecol, 22: 500-503. |
| 28 | Sohrabi M, Rafii M Y, Hanafi M M, Akmar A S N, Latif M A. 2012. Genetic diversity of upland rice germplasm in Malaysia based on quantitative traits.Sci World J, 2012: 1-9. |
| 29 | Song Z P, Xu X, Wang B, Chen J K, Lu B R.2003. Genetic diversity in the northernmost Oryza rufipogon populations estimated by SSR markers.Theor Appl Genet, 107(8): 1492-1499. |
| 30 | Steel R G D, Torrie J H.1980. Principles and Procedures of Statistics. A Biometrical Approach. 2nd edn. New York, USA: Mcgraw-Hill Book Company Inc: 550. |
| 31 | Thomson M J, Septiningsih E M, Suwardjo F, Santoso T J, Silitonga T S, McCouch S R.2007. Genetic diversity analysis of traditional and improved Indonesian rice (Oryza sativa L.) germplasm using microsatellite markers.Theor Appl Genet, 114(3): 559-568. |
| [1] | Veronica Volpe, Franco Magurno, Paola Bonfante, Stefano Ghignone, Erica Lumini. Diversity of Arbuscular Mycorrhizal Fungi Associated with Six Rice Cultivars in Italian Agricultural Ecosystem Managed with Alternate Wetting and Drying [J]. Rice Science, 2023, 30(4): 348-358. |
| [2] | B. M. Lokeshkumar, S. L. Krishnamurthy, Suman Rathor, Arvinder Singh Warriach, N. M. Vinaykumar, B. M. Dushyanthakumar, Parbodh Chander Sharma. Morphophysiological Diversity and Haplotype Analysis of Saltol QTL Region in Diverse Rice Landraces for Salinity Tolerance [J]. Rice Science, 2023, 30(4): 306-320. |
| [3] | Zhou Ying, Wan Tao, Yuan Bin, Lei Fang, Chen Meijuan, Wang Qiong, Huang Ping, Kou Shuyan, Qiu Wenxiu, Liu Li. Improving Rice Blast Resistance by Mining Broad-Spectrum Resistance Genes at Pik Locus [J]. Rice Science, 2022, 29(2): 133-142. |
| [4] | Shalini Pulipati, Suji Somasundaram, Nitika Rana, Kavitha Kumaresan, Mohamed Shafi, Peter Civáň, Gothandapani Sellamuthu, Deepa Jaganathan, Prasanna Venkatesan Ramaravi, S. Punitha, Kalaimani Raju, Shrikant S. Mantri, R. Sowdhamini, Ajay Parida, Gayatri Venkataraman. Diversity of Sodium Transporter HKT1;5 in Genus Oryza [J]. Rice Science, 2022, 29(1): 31-46. |
| [5] | Pathaichindachote Wanwarang, Panyawut Natjaree, Sikaewtung Kannika, Patarapuwadol Sujin, Muangprom Amorntip. Genetic Diversity and Allelic Frequency of Selected Thai and Exotic Rice Germplasm Using SSR Markers [J]. Rice Science, 2019, 26(6): 393-403. |
| [6] | Donde Ravindra, Kumar Jitendra, Gouda Gayatri, Kumar Gupta Manoj, Mukherjee Mitadru, Yasin Baksh Sk, Mahadani Pradosh, Kumar Sahoo Khirod, Behera Lambodar, Kumar Dash Sushanta. Assessment of Genetic Diversity of Drought Tolerant and Susceptible Rice Genotypes Using Microsatellite Markers [J]. Rice Science, 2019, 26(4): 239-247. |
| [7] | F. Aala Jr Wilson, B. Gregorio Glenn. Morphological and Molecular Characterization of Novel Salt-Tolerant Rice Germplasms from the Philippines and Bangladesh [J]. Rice Science, 2019, 26(3): 178-188. |
| [8] | Khlaimongkhon Sudthana, Chakhonkaen Sriprapai, Pitngam Keasinee, Ditthab Khanittha, Sangarwut Numphet, Panyawut Natjaree, Wasinanon Thiwawan, Mongkolsiriwatana Chareerat, Chunwongse Julapark, Muangprom Amorntip. Molecular Markers and Candidate Genes for Thermo-Sensitive Genic Male Sterile in Rice [J]. Rice Science, 2019, 26(3): 147-156. |
| [9] | Wenqiang Liu, Xiaowu Pan, Yongchao Li, Yonghong Duan, Jun Min, Sanxiong Liu, Licheng Liu, Xinnian Sheng, Xiaoxiang Li. Identification of QTLs and Validation of qCd-2 Associated with Grain Cadmium Concentrations in Rice [J]. Rice Science, 2019, 26(1): 42-49. |
| [10] | Kazemi Sheidollah, Reza Eshghizadeh Hamid, Zahedi Morteza. Responses of Four Rice Varieties to Elevated CO2 and Different Salinity Levels [J]. Rice Science, 2018, 25(3): 142-151. |
| [11] | Xiongsiyee Vua, Rerkasem Benjavan, Veeradittakit Jeeraporn, Saenchai Chorpet, Lordkaew Sittichai, Thebault Prom-u-thai Chanakan. Variation in Grain Quality of Upland Rice from Luang Prabang Province, Lao PDR [J]. Rice Science, 2018, 25(2): 94-102. |
| [12] | Radhesh Krishnan Subramanian, Muthuramalingam Pandiyan, Pandian Subramani, Banupriya Ramachandradoss, Chithra Gunasekar, Ramesh Manikandan. Sprouted Sorghum Extract Elicits Coleoptile Emergence, Enhances Shoot and Root Acclimatization, and Maintains Genetic Fidelity in indica Rice [J]. Rice Science, 2018, 25(2): 61-72. |
| [13] | Li Zheng, Chen Liu, Meiling Zhang, Mei Yang, Erxun Zhou. Diversity of dsRNA Viruses Infecting Rice Sheath Blight Fungus Rhizoctonia solani AG-1 IA [J]. Rice Science, 2018, 25(1): 57-60. |
| [14] | Haritha G., P. M. Swamy B., L. Naik M., Jyothi B., Divya B., Malathi S., Sarla N.. Yield Traits and Associated Marker Segregation in Elite Introgression Lines Derived from O. sativa × O. nivara [J]. Rice Science, 2018, 25(1): 19-31. |
| [15] | Naga Bheema Lingeswara Reddy Inja, Kim Sung-Mi, Kim Beom-Ki, Yoon In-Sun, Kwon Taek-Ryoun. Identification of Rice Accessions Associated with K+/Na+ Ratio and Salt Tolerance Based on Physiological and Molecular Responses [J]. Rice Science, 2017, 24(6): 360-364. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||