
Rice Science ›› 2018, Vol. 25 ›› Issue (1): 57-60.DOI: 10.1016/j.rsci.2017.09.002
• Orginal Article • Previous Articles
Li Zheng, Chen Liu, Meiling Zhang, Mei Yang, Erxun Zhou( )
)
Received:2017-07-18
Accepted:2017-09-25
Online:2018-01-28
Published:2017-11-16
Li Zheng, Chen Liu, Meiling Zhang, Mei Yang, Erxun Zhou. Diversity of dsRNA Viruses Infecting Rice Sheath Blight Fungus Rhizoctonia solani AG-1 IA[J]. Rice Science, 2018, 25(1): 57-60.
Add to citation manager EndNote|Ris|BibTeX
| Isolate No. | Location a | Isolate No. | Location a |
|---|---|---|---|
| A5 | Danzhou, Hainan | B315 | Wuhua, Guangdong |
| A90 | Danzhou, Hainan | B316 | Wuhua, Guangdong |
| A92 | Danzhou, Hainan | B331 | Lianzhou, Guangdong |
| A107 | Ledong, Hainan | C40 | Minhou, Fujian |
| A132 | Haikou, Hainan | D164 | Hechi, Guangxi |
| A142 | Haikou, Hainan | D168 | Yangshuo, Guangxi |
| A154 | Wenchang, Hainan | D175 | Yangshuo, Guangxi |
| A182 | Tunchang, Hainan | D181 | Baise, Guangxi |
| B120 | Dongyuan, Guangdong | D186 | Baise, Guangxi |
| B237 | Maoming, Guangdong | E51 | Dali, Yunnan |
| B240 | Maoming, Guangdong | GD-2 | Lechang, Guangdong |
| B246 | Suixi, Guangdong | GD-11 | Ruyuan, Guangdong |
| B249 | Suixi, Guangdong | GD-61 | Guangzhou, Guangdong |
| B250 | Suixi, Guangdong | GD-118 | Doumen, Guangdong |
| B255 | Suixi, Guangdong | H1 | Xuancheng, Anhui |
| B261 | Conghua, Guangdong | H2 | Chaohu, Anhui |
| B265 | Conghua, Guangdong | H3 | Jingzhou, Hunan |
| B266 | Conghua, Guangdong | N1 | Hangzhou, Zhejiang |
| B270 | Xinxing, Guangdong | X-5 | Yangzhou, Jiangsu |
| B275 | Taishan, Guangdong | Y1 | Hangzhou, Zhejiang |
| B278 | Taishan, Guangdong | ZJG-15 | Zhangjiagang, Jiangsu |
| B297 | Lufeng, Guangdong |
Table 1 Details of Rhizoctonia solani AG-1 IA strains used in this study.
| Isolate No. | Location a | Isolate No. | Location a |
|---|---|---|---|
| A5 | Danzhou, Hainan | B315 | Wuhua, Guangdong |
| A90 | Danzhou, Hainan | B316 | Wuhua, Guangdong |
| A92 | Danzhou, Hainan | B331 | Lianzhou, Guangdong |
| A107 | Ledong, Hainan | C40 | Minhou, Fujian |
| A132 | Haikou, Hainan | D164 | Hechi, Guangxi |
| A142 | Haikou, Hainan | D168 | Yangshuo, Guangxi |
| A154 | Wenchang, Hainan | D175 | Yangshuo, Guangxi |
| A182 | Tunchang, Hainan | D181 | Baise, Guangxi |
| B120 | Dongyuan, Guangdong | D186 | Baise, Guangxi |
| B237 | Maoming, Guangdong | E51 | Dali, Yunnan |
| B240 | Maoming, Guangdong | GD-2 | Lechang, Guangdong |
| B246 | Suixi, Guangdong | GD-11 | Ruyuan, Guangdong |
| B249 | Suixi, Guangdong | GD-61 | Guangzhou, Guangdong |
| B250 | Suixi, Guangdong | GD-118 | Doumen, Guangdong |
| B255 | Suixi, Guangdong | H1 | Xuancheng, Anhui |
| B261 | Conghua, Guangdong | H2 | Chaohu, Anhui |
| B265 | Conghua, Guangdong | H3 | Jingzhou, Hunan |
| B266 | Conghua, Guangdong | N1 | Hangzhou, Zhejiang |
| B270 | Xinxing, Guangdong | X-5 | Yangzhou, Jiangsu |
| B275 | Taishan, Guangdong | Y1 | Hangzhou, Zhejiang |
| B278 | Taishan, Guangdong | ZJG-15 | Zhangjiagang, Jiangsu |
| B297 | Lufeng, Guangdong |
| Strain | Location a | dsRNA segment observed | ||
|---|---|---|---|---|
| Number | Size (kb) b | |||
| B275 | Taishan, Guangdong | 2 | 1.7, 2.0 | |
| A5 | Danzhou, Hainan | 3 | 1.7, 1.8, 9.0 | |
| A154 | Wenchang, Hainan | 2 | 1.8, 1.9 | |
| A182 | Tunchang, Hainan | 2 | 1.8, 1.9 | |
| B240 | Maoming, Guangdong | 2 | 1.8, 2.0 | |
| B261 | Conghua, Guangdong | 1 | 1.8 | |
| B266 | Conghua, Guangdong | 4 | 1.8, 2.0, 2.3, 9.0 | |
| B297 | Lufeng, Guangdong | 2 | 1.8, 1.9 | |
| B331 | Lianzhou, Guangdong | 3 | 1.8, 2.0, 2.3 | |
| D168 | Yangshuo, Guangxi | 2 | 1.8, 2.0 | |
| D175 | Yangshuo, Guangxi | 2 | 1.8, 1.9 | |
| GD-11 | Ruyuan, Guangdong | 3 | 1.8, 2.0, 2.3 | |
| H1 | Xuancheng, Anhui | 2 | 1.7, 1.8 | |
| H2 | Chaohu, Anhui | 2 | 1.8, 1.9 | |
| N1 | Hangzhou, Zhejiang | 4 | 1.8, 2.0, 2.3, 6.5 | |
| Y1 | Hangzhou, Zhejiang | 3 | 1.8, 2.0, 6.4 | |
Table 2 Characterization of dsRNA mycoviruses in Rhizoctonia solani AG-1 IA strains.
| Strain | Location a | dsRNA segment observed | ||
|---|---|---|---|---|
| Number | Size (kb) b | |||
| B275 | Taishan, Guangdong | 2 | 1.7, 2.0 | |
| A5 | Danzhou, Hainan | 3 | 1.7, 1.8, 9.0 | |
| A154 | Wenchang, Hainan | 2 | 1.8, 1.9 | |
| A182 | Tunchang, Hainan | 2 | 1.8, 1.9 | |
| B240 | Maoming, Guangdong | 2 | 1.8, 2.0 | |
| B261 | Conghua, Guangdong | 1 | 1.8 | |
| B266 | Conghua, Guangdong | 4 | 1.8, 2.0, 2.3, 9.0 | |
| B297 | Lufeng, Guangdong | 2 | 1.8, 1.9 | |
| B331 | Lianzhou, Guangdong | 3 | 1.8, 2.0, 2.3 | |
| D168 | Yangshuo, Guangxi | 2 | 1.8, 2.0 | |
| D175 | Yangshuo, Guangxi | 2 | 1.8, 1.9 | |
| GD-11 | Ruyuan, Guangdong | 3 | 1.8, 2.0, 2.3 | |
| H1 | Xuancheng, Anhui | 2 | 1.7, 1.8 | |
| H2 | Chaohu, Anhui | 2 | 1.8, 1.9 | |
| N1 | Hangzhou, Zhejiang | 4 | 1.8, 2.0, 2.3, 6.5 | |
| Y1 | Hangzhou, Zhejiang | 3 | 1.8, 2.0, 6.4 | |
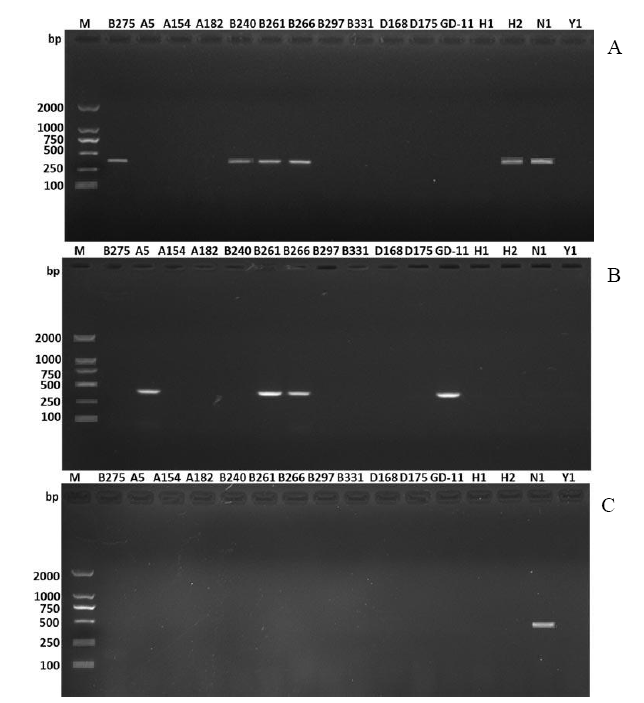
Fig. 2. RT-PCR amplification of three virus species (RsRV1, RsPV2 and RsRV-HN008) in 16 strains of Rhizoctonia solani. The species-specific RT-primers RsRV1F (5′-TGGCGTTGGCTG AGGCAAAGGT-3′) / RsRV1R (5′-GCGCATGATAGTGATGGATACG TGGATTAG-3′), RsPV2F (5′-GTTCAGCGACTACTACGACA-3′) / RsPV2R (5′-CCCCAGAGCATCGGTGTGACGCC-3′) and RsRV- HN008F (5′-GGTGATACGGCTGACAAA-3′) / RsRV-HN008R (5′-ATCG GTTGCCTAACTCCT-3′) designed based on the conserved sequences of RNA dependent RNA polymerase of the three viruses were used to amplify the corresponding conserved fragment of the viruses RsRV1 (A), RsPV2 (B) and RsRV-HN008 (C) in 16 strains of R. solani AG-1 IA, respectively.
| [1] | Bartholomäus A, Wibberg D, Winkler A, Pühler A, Schlüter A, Varrelmann M.2016. Deep sequencing analysis reveals the mycoviral diversity of the virome of an avirulent isolate of Rhizoctonia solani AG-2-2 IV. PLoS One, 11(11): e0165965. |
| [2] | Bharathan N, Saso H, Gudipati L, Bharathan S, Whited K, Anthony K.2005. Double-stranded RNA distribution and analysis among isolates of Rhizoctonia solani AG-2 to -13. Plant Pathol, 54(2): 196-203. |
| [3] | Castanho B, Butler E E.1978. Rhizoctonia decline: A degenerative disease of Rhizoctonia solani. Phytopathology, 68: 1505-1510. |
| [4] | Das S, Falloon R E, Stewart A, Pitman A R.2016. Novel mitoviruses in Rhizoctonia solani AG-3PT infecting potato. Fungal Biol, 120(3): 338-350. |
| [5] | Ghabrial S A.1998. Origin, adapation and evolutionary pathways of fungal viruses.Virus Genes, 16(1): 119-131. |
| [6] | Ghabrial S A, Suzuki N.2008. Fungal viruses. In: Mahy B W J, van Regenmortel M H V. Encyclopedia of Virology. 3rd edn. Oxford: Elsevier: 284-291. |
| [7] | Ghabrial S A, Suzuki N.2009. Viruses of plant pathogenic fungi.Annu Rev Phytopathol, 47: 353-384. |
| [8] | Lin Y H, Chiba S, Tani A, Kondo H, Sasaki A, Kanematsu S, Suzuki N.2012. A novel quadripartite dsRNA virus isolated from a phytopathogenic filamentous fungus,Rosellinia necatrix. Virology, 426(1): 42-50. |
| [9] | Liu L J, Xie J T, Cheng J S, Fu Y P, Li G Q, Yi X H, Jiang D H.2014. Fungal negative-stranded RNA virus that is related to bornaviruses and nyaviruses.Proc Natl Acad Sci USA, 111(33): 12205-12210. |
| [10] | Morris T J, Dodds J A.1979. Isolation and analysis of double stranded RNA from virus infected plant and fungal tissue.Phytopathology, 69(8): 854-858. |
| [11] | Nuss D L.2005. Hypovirulence: Mycoviruses at the fungal-plant interface.Nat Rev Microbiol, 3: 632-642. |
| [12] | Nuss D L.2010. Mycoviruses. In: Borkovich K A. Cellular and Molecular Biology of Filamentous Fungi. Washington, USA: American Society of Microbiology: 145-152. |
| [13] | Ogoshi A.1987. Ecology and pathogenicity of anastomosis and intraspecific groups of Rhizoctonia solani Kühn. Annu Rev Phytopathol, 25: 125-143. |
| [14] | Okada R, Kiyota E, Sabanadzovic S, Moriyama H, Fukuhara T, Saha P, Roossinck M J, Severin A, Valverde R A.2011. Bell pepper endornavirus: Molecular and biological properties, and occurrence in the genus Capsicum. J Gen Virol, 92: 2664-2673. |
| [15] | Pearson M N, Beever R E, Boine B, Arthur K.2009. Mycoviruses of filamentous fungi and their relevance to plant pathology.Mol Plant Pathol, 10(1): 115-128. |
| [16] | Vainio E J, Hyder R, Aday G, Hansen E, Piri T, Dogmus-Lehtijarvi T, Lehtijarvi A, Korhonen K, Hantula J.2012. Population structure of a novel putative mycovirus infecting the conifer root-rot fungus Heterobasidium annosum sensu lato. Virology, 422(2): 366-376. |
| [17] | Yu X, Li B, Fu Y P, Jiang D H, Ghabrial S A, Li G Q, Peng Y L, Xie J T, Cheng J S, Huang J, Yi X H.2010. A geminivirus-related DNA mycovirus that confers hypovirulence to a plant pathogenic fungus.Proc Natl Acad Sci USA, 107(18): 8387-8392. |
| [18] | Yu X, Li B, Fu Y P, Xie J T, Cheng J S, Ghabrial S A, Li G Q, Yi X H, Jiang D H.2013. Extracellular transmission of a DNA mycovirus and its use as a nature fungicide.Proc Natl Acad Sci USA, 110(4): 1452-1457. |
| [19] | Zhang D T, Peng Z K, Cao Q Q, Yang M, Zheng L, Zhou E X.2012. Colonization of three antagonistic strains on rice plant and their biocontrol effects on rice sheath blight.J Northwest Agric Forest Univ, 40: 97-102. (in Chinese with English abstract) |
| [20] | Zheng L, Liu H Q, Zhang M L, Cao X, Zhou E X.2013. The complete genomic sequence of a novel mycovirus from Rhizoctonia solani AG-1 IA strain B275. Arch Virol, 158(7): 1609-1612. |
| [21] | Zheng L, Zhang M L, Chen Q G, Zhu M H, Zhou E X.2014. A novel mycovirus closely related to viruses in the genus Alphapartitivirus confers hypovirulence in the phytopathogenic fungus Rhizoctonia solani. Virology, 456/457(4): 220-226. |
| [22] | Zhong J, Chen C Y, Gao B D.2015. Genome sequence of a novel mycovirus of Rhizoctonia solani, a plant pathogenic fungus. Virus Genes, 51(1): 167-170. |
| [1] | Veronica Volpe, Franco Magurno, Paola Bonfante, Stefano Ghignone, Erica Lumini. Diversity of Arbuscular Mycorrhizal Fungi Associated with Six Rice Cultivars in Italian Agricultural Ecosystem Managed with Alternate Wetting and Drying [J]. Rice Science, 2023, 30(4): 348-358. |
| [2] | B. M. Lokeshkumar, S. L. Krishnamurthy, Suman Rathor, Arvinder Singh Warriach, N. M. Vinaykumar, B. M. Dushyanthakumar, Parbodh Chander Sharma. Morphophysiological Diversity and Haplotype Analysis of Saltol QTL Region in Diverse Rice Landraces for Salinity Tolerance [J]. Rice Science, 2023, 30(4): 306-320. |
| [3] | Wang Chenjiaozi, Zhao Mei, Shu Canwei, Zhou Erxun. Three Genes Related to Trehalose Metabolism Affect Sclerotial Development of Rhizoctonia solani AG-1 IA, Causal Agent of Rice Sheath Blight [J]. Rice Science, 2022, 29(3): 268-276. |
| [4] | Zhou Ying, Wan Tao, Yuan Bin, Lei Fang, Chen Meijuan, Wang Qiong, Huang Ping, Kou Shuyan, Qiu Wenxiu, Liu Li. Improving Rice Blast Resistance by Mining Broad-Spectrum Resistance Genes at Pik Locus [J]. Rice Science, 2022, 29(2): 133-142. |
| [5] | Shalini Pulipati, Suji Somasundaram, Nitika Rana, Kavitha Kumaresan, Mohamed Shafi, Peter Civáň, Gothandapani Sellamuthu, Deepa Jaganathan, Prasanna Venkatesan Ramaravi, S. Punitha, Kalaimani Raju, Shrikant S. Mantri, R. Sowdhamini, Ajay Parida, Gayatri Venkataraman. Diversity of Sodium Transporter HKT1;5 in Genus Oryza [J]. Rice Science, 2022, 29(1): 31-46. |
| [6] | Pathaichindachote Wanwarang, Panyawut Natjaree, Sikaewtung Kannika, Patarapuwadol Sujin, Muangprom Amorntip. Genetic Diversity and Allelic Frequency of Selected Thai and Exotic Rice Germplasm Using SSR Markers [J]. Rice Science, 2019, 26(6): 393-403. |
| [7] | Donde Ravindra, Kumar Jitendra, Gouda Gayatri, Kumar Gupta Manoj, Mukherjee Mitadru, Yasin Baksh Sk, Mahadani Pradosh, Kumar Sahoo Khirod, Behera Lambodar, Kumar Dash Sushanta. Assessment of Genetic Diversity of Drought Tolerant and Susceptible Rice Genotypes Using Microsatellite Markers [J]. Rice Science, 2019, 26(4): 239-247. |
| [8] | Kazemi Sheidollah, Reza Eshghizadeh Hamid, Zahedi Morteza. Responses of Four Rice Varieties to Elevated CO2 and Different Salinity Levels [J]. Rice Science, 2018, 25(3): 142-151. |
| [9] | Xiongsiyee Vua, Rerkasem Benjavan, Veeradittakit Jeeraporn, Saenchai Chorpet, Lordkaew Sittichai, Thebault Prom-u-thai Chanakan. Variation in Grain Quality of Upland Rice from Luang Prabang Province, Lao PDR [J]. Rice Science, 2018, 25(2): 94-102. |
| [10] | Saakre Manjesh, Meera Baburao Thirthikar, Puthenpeedikal Salim Abida, Mary Ffancies Rose, Poothecty Achuthan Valasala, Thomas George, Radha Sivarajan Sajeevan. Identification and Characterization of Genes Responsible for Drought Tolerance in Rice Mediated by Pseudomonas fluorescens [J]. Rice Science, 2017, 24(5): 291-298. |
| [11] | Rekha Talukdar Preeti, Rathi Sunayana, Pathak Khanin, Kumar Chetia Sanjay, Nath Sarma Ramendra. Population Structure and Marker-Trait Association in Indigenous Aromatic Rice [J]. Rice Science, 2017, 24(3): 145-154. |
| [12] | Anupam Alpana, Imam Jahangir, Mohammad Quatadah Syed, Siddaiah Anantha, Prasad Das Shankar, Variar Mukund, Prasad Mandal Nimai. Genetic Diversity Analysis of Rice Germplasm in Tripura State of Northeast India Using Drought and Blast Linked Markers [J]. Rice Science, 2017, 24(1): 10-20. |
| [13] | Haritha G., Sudhakar T., Chandra D., Ram T., Divya B., Sarla N.. Informative ISSR Markers Help Identify Genetically Distinct Accessions of Oryza rufipogon in Yield Improvement [J]. Rice Science, 2016, 23(5): 225-241. |
| [14] | Sharf Uddin Ahmed Mir, Khalequzzaman Mohammad, Khairul Bashar Md., Kalam Mohammad Shamsuddin Abul. Agro-Morphological, Physico-Chemical and Molecular Characterization of Rice Germplasm with Similar Names of Bangladesh [J]. Rice Science, 2016, 23(4): 211-218. |
| [15] | Rahman Moni Zakiah, Ansar Ali Md, Shahidul Alam Md, Asif Rahman Md, Rejwan Bhuiyan Md, Salim Mian Md, Md Iftekharuddaula Khandakar, Abdul Latif Md, Ashik Iqbal Khan Mohammad. Morphological and Genetical Variability among Rhizoctonia solani Isolates Causing Sheath Blight Disease of Rice [J]. Rice Science, 2016, 23(1): 42-50. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||