
Rice Science ›› 2017, Vol. 24 ›› Issue (4): 198-206.DOI: 10.1016/j.rsci.2017.01.002
• Orginal Article • Previous Articles Next Articles
S. M. Masuduzzaman A.1, Maksudul Haque Md.2( ), K. M. Shamsuddin A.3, A. Salam M.1, Ansar Ali Md.4
), K. M. Shamsuddin A.3, A. Salam M.1, Ansar Ali Md.4
Received:2016-10-06
Accepted:2017-01-13
Online:2017-07-10
Published:2017-04-28
S. M. Masuduzzaman A., Maksudul Haque Md., K. M. Shamsuddin A., A. Salam M., Ansar Ali Md.. Haplotype Diversity at Sub1 Locus and Allelic Distribution Among Rice Varieties of Tide and Flood Prone Areas of South-East Asia[J]. Rice Science, 2017, 24(4): 198-206.
Add to citation manager EndNote|Ris|BibTeX
| Sub1A locus | Sub1C locus | |||||||
|---|---|---|---|---|---|---|---|---|
| Sub1A0 | Sub1A1 | Sub1A2 | Sub1C1 | Sub1C2 | Sub1C3 | Sub1C4 | ||
| Allele size before digestion (bp) | - | 241 | 241 | 175 | 165 | 170 | 175 | |
| Allele size after digestion (bp) | - | 241 | 134, 107 | 130, 45 | 110, 55 | 110, 60 | 110, 65 | |
| Account | 89 | 16 | 55 | 12 | 83 | 6 | 59 | |
| Frequency (%) | 55.6 | 10 | 34.4 | 7.5 | 51.9 | 3.7 | 36.9 | |
Table 1 Size and frequency of different variants of Sub1A and Sub1C alleles, before and after digestion with respective enzymes.
| Sub1A locus | Sub1C locus | |||||||
|---|---|---|---|---|---|---|---|---|
| Sub1A0 | Sub1A1 | Sub1A2 | Sub1C1 | Sub1C2 | Sub1C3 | Sub1C4 | ||
| Allele size before digestion (bp) | - | 241 | 241 | 175 | 165 | 170 | 175 | |
| Allele size after digestion (bp) | - | 241 | 134, 107 | 130, 45 | 110, 55 | 110, 60 | 110, 65 | |
| Account | 89 | 16 | 55 | 12 | 83 | 6 | 59 | |
| Frequency (%) | 55.6 | 10 | 34.4 | 7.5 | 51.9 | 3.7 | 36.9 | |
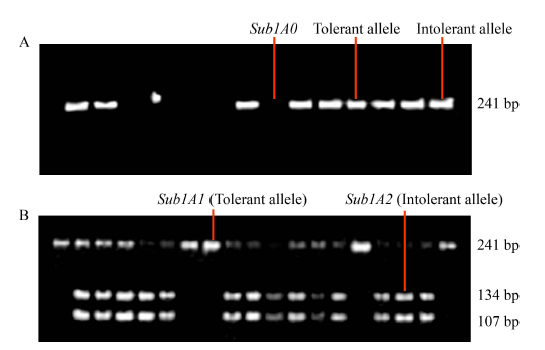
Fig. 1. Sub1A0, Sub1A1 and Sub1A2 alleles among varieties. A, Single allele before digestion with Alu I; B, Two allelic variants after digestion with Alu I.IR40931 and IR42 have tolerant and intolerant allele, respectively.
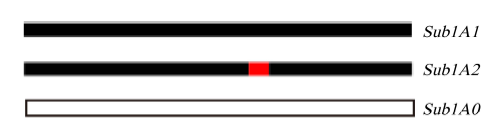
Fig. 2. Allelic patterns generated at Sub1A.Sub1A1, Fragment without restriction site; Sub1A2, Red mark indicates a single nucleotide polymorphism at a specific site in the DNA sequence; Sub1A0, A null allele that completely lacks the gene’s normal function.
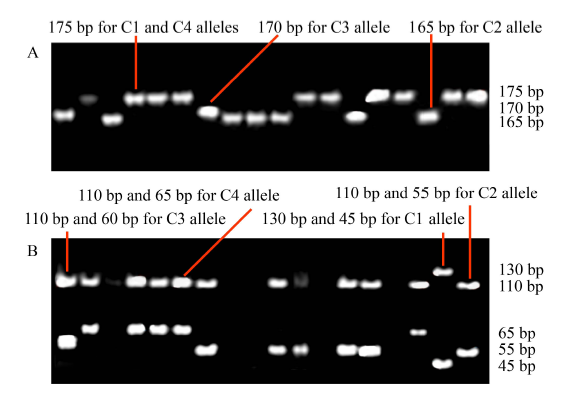
Fig. 3. Sub1C alleles among varieties. A, Three allelic variants before digestion with Cac8 I; B, Four allelic variants after digestion with Cac8 I. IR40931, IR42, Lemo, Matichak have Sub1C1, Sub1C2, Sub1C3, and Sub1C4 alleles, respectively. Sub1C1, 175 bp is cut into 130 and 45 bp (tolerant specific); Sub1C4, 175 bp is cut into 110 and 65 bp (intolerant specific); Sub1C3, 170 bp is cut into 110 and 60 bp (intolerant specific); Sub1C2, 165 bp is cut into 110 and 55 bp (intolerant specific).
| Country/Institute | A1C1 | A1C2 | A1C4 | A2C2 | A2C4 | A0C2 | A0C3 | A0C4 | Total |
|---|---|---|---|---|---|---|---|---|---|
| Bangladesh | 2 | 2 | 1 | 22 | 0 | 15 | 1 | 7 | 50 |
| India | 2 | 1 | 0 | 4 | 1 | 2 | 0 | 4 | 14 |
| Sri Lanka | 7 | 0 | 0 | 6 | 0 | 0 | 1 | 2 | 16 |
| Indonesia | 0 | 0 | 0 | 19 | 1 | 9 | 3 | 37 | 69 |
| Vietnam | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 5 | 7 |
| IRRI | 1 | 0 | 0 | 2 | 0 | 0 | 0 | 1 | 4 |
| Total | 12 | 3 | 1 | 53 | 2 | 27 | 6 | 56 | 160 |
| Frequency (%) | 7.5 | 1.9 | 0.6 | 33.1 | 1.3 | 16.9 | 3.7 | 35 | 100 |
Table 2 Pattern of haplotype frequencies and their distributions.
| Country/Institute | A1C1 | A1C2 | A1C4 | A2C2 | A2C4 | A0C2 | A0C3 | A0C4 | Total |
|---|---|---|---|---|---|---|---|---|---|
| Bangladesh | 2 | 2 | 1 | 22 | 0 | 15 | 1 | 7 | 50 |
| India | 2 | 1 | 0 | 4 | 1 | 2 | 0 | 4 | 14 |
| Sri Lanka | 7 | 0 | 0 | 6 | 0 | 0 | 1 | 2 | 16 |
| Indonesia | 0 | 0 | 0 | 19 | 1 | 9 | 3 | 37 | 69 |
| Vietnam | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 5 | 7 |
| IRRI | 1 | 0 | 0 | 2 | 0 | 0 | 0 | 1 | 4 |
| Total | 12 | 3 | 1 | 53 | 2 | 27 | 6 | 56 | 160 |
| Frequency (%) | 7.5 | 1.9 | 0.6 | 33.1 | 1.3 | 16.9 | 3.7 | 35 | 100 |

Fig. 4. Allelic patterns generated at Sub1C after digestion with Cac8 I.Restriction region is red at different positions. After digestion with Cac8 I, four allelic variants were found.
| Haplotype | Representative variety | Phenotype | Average of elongation score | Average survival rate (%) | Elongation rate (%) |
|---|---|---|---|---|---|
| A1C1 | FR13A | SE/T | 2.3 | 94 | 37 |
| Motorsail | ME/MT | 5.1 | 59 | 78 | |
| Gopalbhag | HE/MS | 7.3 | 36 | 93 | |
| A2C2 | Madabaru Kottamali | ME/MT | 3 | 74 | 56 |
| Habiganj Aman 7 | HE/S | 7.7 | 18 | 212 | |
| A0C2 | Badal | ME/MT | 4.3 | 52 | 64 |
| Fulkari | HE/S | 7.3 | 0 | 167 | |
| A0C4 | Kannimurunga | ME/MT | 5.3 | 63 | 72 |
| Kumragoir | HE/S | 9 | 17 | 123 | |
| A1C2 | Lalchikon | SE/MT | 4.5 | 58 | 47 |
| A1C4 | Matichak | ME/MT | 5 | 60 | 80 |
| A2C4 | CN540 | HE/S | 9 | 26 | 102 |
| A0C3 | Ketumbar | HE/S | 8.7 | 7 | 110 |
Table 3 Association of haplotype patterns with phenotype (elongation and survival).
| Haplotype | Representative variety | Phenotype | Average of elongation score | Average survival rate (%) | Elongation rate (%) |
|---|---|---|---|---|---|
| A1C1 | FR13A | SE/T | 2.3 | 94 | 37 |
| Motorsail | ME/MT | 5.1 | 59 | 78 | |
| Gopalbhag | HE/MS | 7.3 | 36 | 93 | |
| A2C2 | Madabaru Kottamali | ME/MT | 3 | 74 | 56 |
| Habiganj Aman 7 | HE/S | 7.7 | 18 | 212 | |
| A0C2 | Badal | ME/MT | 4.3 | 52 | 64 |
| Fulkari | HE/S | 7.3 | 0 | 167 | |
| A0C4 | Kannimurunga | ME/MT | 5.3 | 63 | 72 |
| Kumragoir | HE/S | 9 | 17 | 123 | |
| A1C2 | Lalchikon | SE/MT | 4.5 | 58 | 47 |
| A1C4 | Matichak | ME/MT | 5 | 60 | 80 |
| A2C4 | CN540 | HE/S | 9 | 26 | 102 |
| A0C3 | Ketumbar | HE/S | 8.7 | 7 | 110 |

Fig. 5. Recovery of varieties after submergence.IR40931 is as the tolerant check (A1C1), Madabaru is as the moderately tolerant check (A2C2) and IR42 is as the intolerant check (A2C2). A, 7 d after de-submergence; B, 14 d after de-submergence; C, 40 d after de-submergence.
| 1 | Ahmed N, Siddiqui A A, Ahmed A.2000. DDRT-PCR: Use of agarose gels for detection of amplified products. Mol Vis, 6(19): 144-147. |
| 2 | Bai G H, Guo P G, Kolb F L.2003. Genetic relationships among head blight resistant cultivars of wheat assessed on the basis of molecular markers.Crop Sci, 43(2): 498-507. |
| 3 | Bailey-Serres J, Fukao T, Ronald P, Ismail A, Heuer S, Mackill D.2010. Submergence tolerant rice: SUB1’s journey from landrace to modern cultivar.Rice, 3(2): 138-147. |
| 4 | Chapman J M, Cooper J D, Todd J A, Clayton D G.2003. Detecting disease associations due to linkage disequilibrium using haplotype tags: A class of tests and the determinants of statistical power.Hum Hered, 56: 18-31. |
| 5 | Chinq A, Caldwell K S, Junq M, Dolan M, Smith O S, Tinger S, Morgante M, Rafalski A J.2002. SNP frequency, haplotype structure and linkage disequilibrium in elite maize inbred lines.BMC Genet, 3: 19. |
| 6 | Clark A G, Weiss K M, Nickerson D A, Taylor S L, Buchanan A, Stengard J, Salomaa V, Vartiainen E, Perola M, Boerwinkle E, Sing C F.1998. Haplotype structure and population genetic inferences from nucleotide-sequence variation in human lipoprotein lipase.Am J Hum Genet, 63(2): 595-612. |
| 7 | Clark T G, Andrew T, Cooper G M, Margulies E H, Mullikin J C, Balding D J.2007. Functional constraint and small insertions and deletions in the ENCODE regions of the human genome.Genome Biol, 8: 180. |
| 8 | Collard B C Y, Septiningsih E M, Das S R, Carandang J J, Pamplona A M, Sanchez D L, Kato Y, Ye G, Reddy J N, Singh U S, Iftekharuddaula K M, Venuprasad R, Vera-Cruz C N, Mackill D J, Ismail A M.2013. Developing new flood-tolerant varieties at the International Rice Research Institute (IRRI).SABRAO J Breeding Genet, 45(1): 42-56. |
| 9 | Daly M J, Rioux J D, Schaffner S F, Hudson T J, Lander E S.2001. High-resolution haplotype structure in the human genome.Nat Genet, 29: 229-232. |
| 10 | Ella E S, Kawano N, Yamauchi Y, Tanaka K, Ismail A M.2003. Blocking ethylene perception enhances flooding tolerance in rice.Func Plant Biol, 30(7): 813-819. |
| 11 | Estoup A, Tailliez C, Cornuet J M, Solignac M.1995. Size homoplasy and mutational processes of interrupted microsatellites in two bee species, Apis mellifera and Bombus terrestris (Apidae).Mol Biol Evol, 12(6): 1074-1084. |
| 12 | Glaubitz J C, Rhodes O E, Dewoody J A.2003. Prospects for inferring pairwise relationships with single nucleotide polymorphisms.Mol Ecol, 12(4): 1039-1047. |
| 13 | Gupta P K, Roy J K, Prasad M.2001. Single nucleotide polymorphisms: A new paradigm for molecular marker technology and DNA polymorphism detection with emphasis on their use in plants.Curr Sci, 80(4): 524-535. |
| 14 | Hampe J, Schreiber S, Krawczak M.2003. Entropy-based SNP selection for genetic association studies.Hum Genet, 114(1): 36-43. |
| 15 | Iftekharuddaula K M, Newaz M A, Salam M A, Ahmed H U, Mahbub M A A, Septiningsih E M, Collard B C Y, Sanchez D L, Pamplona A M, Mackill D J.2011. Rapid and highprecision marker-assisted backcrossing to introgress the SUB1 QTL into BR11, the rainfed lowland rice mega variety of Bangladesh.Euphytica, 178(1): 83-97. |
| 16 | Jackson M B, Waters I, Setter T, Greenway H.1987. Injury to rice plants caused by complete submergence: A contribution of ethylene.J Exp Bot, 38(11): 1826-1838. |
| 17 | Jackson M B, Ram P C.2003. Physiological and molecular basis of susceptibility and tolerance of rice plants to complete submergence.Ann Bot, 91(2): 227-241. |
| 18 | Johnson G C L, Esposito L, Barratt B J, Smith A N, Heward J, Genova G D, Ueda H, Cordell H J, Eaves I A, Dudbridge F, Twells R C J, Payne F, Hughes W, Nutland S, Stevens H, Carr P, Tuomilehto-Wolf E, Tuomilehto J, Gough S C L, Clayton D G, Todd J A.2001. Haplotype tagging for the identification of common disease genes.Nat Genet, 29: 233-237. |
| 19 | Judson R, Salisbury B, Schneider J, Windemuth A, Stephens J C.2002. How many SNPs does a genome-wide haplotype map require?Pharmacogenomics, 3(3): 379-391. |
| 20 | Kruglyak L.1999. Prospects for whole-genome linkage disequilibrium mapping of common disease genes.Nat Genet, 22: 139-144. |
| 21 | Liu J F, Papasian C, Deng H W.2007. Incorporating single-locus tests into haplotype cladistic analysis in case-control studies.PLoS Genet, 3(3): e46. |
| 22 | Lorenz T C.2012. Polymerase chain reaction: Basic protocol plus troubleshooting and optimization strategies.J Vis Exp, 63: e3998. |
| 23 | Lu Y L, Shah T, Hao Z F, Taba S, Zhang S H, Gao S B, Liu J, Cao M J, Wang J, Prakash A B, Rong T Z, Xu Y B.2011. Comparative SNP and haplotype analysis reveals a higher genetic diversity and rapider LD decay in tropical than temperate germplasm in maize.PLoS One, 6(9): e24861. |
| 24 | Luca M D, Roshina N V, Geiger-Thornsberry G L, Lyman R F, Pasyukova E G, Mackay T F C.2003. Dopa decarboxylase (Ddc) affects variation in drosophila longevity.Nat Genet, 34: 429-433. |
| 25 | Mackill D J.1986. Rainfed lowland rice improvement in South and Southeast Asia: Results of a survey. In: Progress in Rainfed Lowland Rice. Los Baños, the Philippines: International Rice Research Institute: 115-144. |
| 26 | McCartney C A, Somers D J, Fedak G, Cao W.2004. Haploytpe diversity at fusarium head blight resistance QTLs in wheat.Theor Appl Genet, 109(2): 261-271. |
| 27 | McLauchlan A, Ogbonnaya F C, Hollingsworth B, Carter M, Gale K R, Henry R J, Holton T A, Morell M K, Rampling L P, Sharp P J, Shariflou M R, Jones M G K, Appels R.2001. Development of robust PCR-based DNA markers for each homoeo-allele of granule-bound starch synthase and their application in wheat breeding programs.Aust J Agric Res, 52(11/12): 1409-1416. |
| 28 | Mispan M S, Zhang L, Feng J, Gu X Y.2013. Quantitative trait locus and haplotype analyses of wild and crop-mimic traits in U.S. weedy rice.G3, 3(6): 1049-1059. |
| 29 | Mogg R, Batley J, Hanley S, Edwards D, O’Sullivan H, Edwards K.2002. Characterising the flanking regions of Zea mays microsatellites reveals a large number of useful sequence polymorphisms.Theor Appl Genet, 105(4): 532-543. |
| 30 | Neeraja C N, Maghirang-Rodriguez R, Pamplona A, Heuer S, Collard B C Y, Septiningsih E M, Vergara G, Sanchez D, Xu K, Ismail A M, Mackill D J.2007. A marker-assisted backcross approach for developing submergence-tolerant rice cultivars.Theor Appl Genet, 115(6): 767-776. |
| 31 | Rafalski A.2002. Applications of single nucleotide polymorphisms in crop genetics.Curr Opin Plant Biol, 5(2): 94-100. |
| 32 | Reed D H, Frankham R.2003. Correlation between fitness and genetic diversity regulatory variation in mouse genes.Nat Genet, 32: 1532-1739. |
| 33 | Sardana S.1997. Submergence tolerance and elongation ability in newly developed lines of rice for Tripura.Ind J Hill Farming, 10(1/2): 111-112. |
| 34 | Sasaki R Z C, Zhao Z C.2000a. Submergence tolerance of transplanted rice seedlings with several plant ages in leaf number: The duration of submergence and the role of endosperm reserves.Jpn J Crop Sci, 69(3): 365-371. |
| 35 | Sasaki R Z C, Zhao Z C.2000b. Submergence tolerance of transplanted rice seedlings with several plant ages in leaf number: Relationship between the growth during submergence and survival.Jpn J Crop Sci, 69(3): 372-379. |
| 36 | Septiningsih E M, Pamplona A M, Sanchez D L, Neeraja C N, Vergara G V, Heuer S, Ismail A M, Mackill D J.2009. Development of submergence-tolerant rice cultivars: The Sub1 locus and beyond.Ann Bot, 103(2): 151-160. |
| 37 | Septiningsih E M, Collard B C Y, Heuer S, Bailey-Serres J, Ismail A M, Mackill D J.2013. Applying genomics tools for breeding submergence tolerance in rice. In: Varshney R K, Tuberosa R. Translational Genomics for Crop Breeding: Improvement for Abiotic Stress, Quality and Yield Improvement. USA: Wiley- Blackwell: 9-30. |
| 38 | Septiningsih E M, Hidayatun N, Sanchez D L, Nugraha Y, Carandang J, Pamplona A M, Collard B C Y, Ismail A M, Mackill D J.2015. Accelerating the development of new submergence tolerant rice varieties: The case of Ciherang-Sub1 and PSB Rc18-Sub1.Euphytica, 202(2): 259-268. |
| 39 | Servin B, Martin O C, Me´zard M, Hospital F.2004. Toward a theory of marker-assisted gene pyramiding.Genetics, 168(1): 513-523. |
| 40 | Setter T L, Ellis M, Laureles E V, Ella E S, Senadhira D, Mishra S B, Sarkarung S, Datta S.1997. Physiology and genetics of submergence tolerance in rice.Ann Bot, 79: 67-77. |
| 41 | Shariflou M R, Sharp P J.1999. A polymorphic microsatellite in the 3’ end of ‘waxy’ genes of wheat, Triticum aestivum.Plant Breeding, 118(3): 275-277. |
| 42 | Singh N, Dang T T M, Vergara G V, Pandey D M, Sanchez D, Neeraja C N, Septiningsih E M, Mendioro M, Tecson-Mendoza E M, Ismail A M, Mackill D J, Heuer S.2010. Molecular marker survey and expression analyses of the rice submergence tolerance gene SUB1A.Theor Appl Genet, 121(8): 1441-1453. |
| 43 | Stephens J C, Schneider J A, Tanguay D A, Choi J, Acharya T, Stanley S E, Jiang R, Messer C J, Chew A, Han J H, Duan J C, Carr J L, Lee M S, Koshy B, Kumar A M, Zhang G, Newell W R, Windemuth A, Xu C B, Kalbfleisch T S, Shaner S L, Arnold K, Schulz V, Drysdale C M, Nandabalan K, Judson R S, Ruano G, Vovis G F.2001. Haplotype variation and linkage disequilibrium in 313 human genes.Science, 293: 489-493. |
| 44 | Viard F, Franck P, Dubois M P, Estoup A, Jarne P.1998. Variation of microsatellite size homoplasy across electromorphs, loci, and populations in three invertebrate species.J Mol Evol, 47(1): 42-51. |
| 45 | Wang Z K, Ni Z F, Wu H L, Nie X L, Sun Q X.2006. Heterosis in root development and differential gene expression between hybrids and their parental inbreds in wheat (Triticum aestivum L.).Theor Appl Genet, 113(7): 1283-1294. |
| 46 | Xu K, Mackill D J.1996. A major locus for submergence tolerance mapped on rice chromosome 9.Mol Breeding, 2(3): 219-224. |
| 47 | Xu K, Xu X, Ronald P C, Mackill D J.2000. A high-resolution linkage map of the vicinity of the rice submergence tolerance locus Sub1.Mol Gen Genet, 263(4): 681-689. |
| 48 | Xu K, Xu X, Fukao T, Canlas P, Maghirang-Rodriguez R, Heuer S, Ismail A M, Bailey-Serres J, Ronald P C, Mackill D J.2006. Sub1A is an ethylene-response-factor-like gene that confers submergence tolerance to rice.Nature, 442: 705-708. |
| 49 | Yamada N.1959. Physiological basis of resistance of rice plant against overhead flooding.Bull Nat Inst Agric Sci, 8: 110. |
| 50 | Yamamori M, Quynh N T.2000. Differential effects of Wx-A1, -B1 and -D1 protein deficiencies on apparent amylose content and starch pasting properties in common wheat.Theor Appl Genet, 100(1): 32-38. |
| 51 | Zhang K, Sun F Z, Zhao H Y.2005. Haplore: A program for haplotype reconstruction in general pedigrees without recombination.Bioinformatics, 21(1): 90-103. |
| [1] | B. M. Lokeshkumar, S. L. Krishnamurthy, Suman Rathor, Arvinder Singh Warriach, N. M. Vinaykumar, B. M. Dushyanthakumar, Parbodh Chander Sharma. Morphophysiological Diversity and Haplotype Analysis of Saltol QTL Region in Diverse Rice Landraces for Salinity Tolerance [J]. Rice Science, 2023, 30(4): 306-320. |
| [2] | Liu Yantong, Li Ting, Jiang Zhishu, Zeng Chuihai, He Rong, Qiu Jiao, Lin Xiaoli, Peng Limei, Song Yongping, Zhou Dahu, Cai Yicong, Zhu Changlan, Fu Junru, He Haohua, Xu Jie. Characterization of a Novel Weak Allele of RGA1/D1 and Its Potential Application in Rice Breeding [J]. Rice Science, 2022, 29(6): 522-534. |
| [3] | Zhou Ying, Wan Tao, Yuan Bin, Lei Fang, Chen Meijuan, Wang Qiong, Huang Ping, Kou Shuyan, Qiu Wenxiu, Liu Li. Improving Rice Blast Resistance by Mining Broad-Spectrum Resistance Genes at Pik Locus [J]. Rice Science, 2022, 29(2): 133-142. |
| [4] | Shalini Pulipati, Suji Somasundaram, Nitika Rana, Kavitha Kumaresan, Mohamed Shafi, Peter Civáň, Gothandapani Sellamuthu, Deepa Jaganathan, Prasanna Venkatesan Ramaravi, S. Punitha, Kalaimani Raju, Shrikant S. Mantri, R. Sowdhamini, Ajay Parida, Gayatri Venkataraman. Diversity of Sodium Transporter HKT1;5 in Genus Oryza [J]. Rice Science, 2022, 29(1): 31-46. |
| [5] | Yanchang Luo, Tingchen Ma, Teo Joanne, Zhixiang Luo, Zefu Li, Jianbo Yang, Zhongchao Yin. Marker-Assisted Breeding of Thermo-Sensitive Genic Male Sterile Line 1892S for Disease Resistance and Submergence Tolerance [J]. Rice Science, 2021, 28(1): 89-98. |
| [6] | Panda Debabrata, Barik Jijnasa. Flooding Tolerance in Rice: Focus on Mechanisms and Approaches [J]. Rice Science, 2021, 28(1): 43-57. |
| [7] | Barik Jijnasa, Kumar Vajinder, K. Lenka Sangram, Panda Debabrata. Assessment of Variation in Morpho-Physiological Traits and Genetic Diversity in Relation to Submergence Tolerance of Five Indigenous Lowland Rice Landraces [J]. Rice Science, 2020, 27(1): 32-43. |
| [8] | Yang Lv, Yueying Wang, Jahan Noushin, Haitao Hu, Ping Chen, Lianguang Shang, Haiyan Lin, Guojun Dong, Jiang Hu, Zhenyu Gao, Qian Qian, Yu Zhang, Longbiao Guo. Genome-Wide Association Analysis and Allelic Mining of Grain Shape-Related Traits in Rice [J]. Rice Science, 2019, 26(6): 384-392. |
| [9] | Donde Ravindra, Kumar Jitendra, Gouda Gayatri, Kumar Gupta Manoj, Mukherjee Mitadru, Yasin Baksh Sk, Mahadani Pradosh, Kumar Sahoo Khirod, Behera Lambodar, Kumar Dash Sushanta. Assessment of Genetic Diversity of Drought Tolerant and Susceptible Rice Genotypes Using Microsatellite Markers [J]. Rice Science, 2019, 26(4): 239-247. |
| [10] | T. Bui Liem, S. Ella Evangelina, L. Dionisio-Sese Maribel, M. Ismail Abdelbagi. Morpho-Physiological Changes in Roots of Rice Seedling upon Submergence [J]. Rice Science, 2019, 26(3): 167-177. |
| [11] | Yaobin Qin, Peng Cheng, Yichen Cheng, Yue Feng, Derun Huang, Tingxu Huang, Xianjun Song, Jiezheng Ying. QTL-Seq Identified a Major QTL for Grain Length and Weight in Rice Using Near Isogenic F2 Population [J]. Rice Science, 2018, 25(3): 121-131. |
| [12] | Chandra Roy Subhas, Bhasker Reddy Lachagari Vijaya. Assessment of SNP and InDel Variations Among Rice Lines of Tulaipanji x Ranjit [J]. Rice Science, 2017, 24(6): 336-348. |
| [13] | Srivastava Deepti, Shamim Md, Kumar Mahesh, Mishra Anurag, Pandey Pramila, Kumar Deepak, Yadav Prashant, Harrish Siddiqui Mohammed, Narayan Singh Kapildeo. Current Status of Conventional and Molecular Interventions for Blast Resistance in Rice [J]. Rice Science, 2017, 24(6): 299-321. |
| [14] | D. Chowdhury A., Haritha G., Sunitha T., L. Krishnamurthy S., Divya B., Padmavathi G., Ram T., Sarla N.. Haplotyping of Rice Genotypes Using Simple Sequence Repeat Markers Associated with Salt Tolerance [J]. Rice Science, 2016, 23(6): 317-325. |
| [15] | Ya-fang Zhang, Yu-yin Ma, Zong-xiang Chen, Jie Zou, Tian-xiao Chen, Qian-qian Li, Xue-biao Pan, Shi-min Zuo. Genome-Wide Association Studies Reveal New Genetic Targets for Five Panicle Traits of International Rice Varieties [J]. Rice Science, 2015, 22(5): 217-226. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||