
Rice Science ›› 2018, Vol. 25 ›› Issue (2): 73-81.DOI: 10.1016/j.rsci.2017.11.002
• Orginal Article • Previous Articles Next Articles
Nurdiani Dini, Widyajayantie Dwi, Nugroho Satya( )
)
Received:2017-08-04
Accepted:2017-11-06
Online:2018-03-28
Published:2017-12-22
Nurdiani Dini, Widyajayantie Dwi, Nugroho Satya. OsSCE1 Encoding SUMO E2-Conjugating Enzyme Involves in Drought Stress Response of Oryza sativa[J]. Rice Science, 2018, 25(2): 73-81.
Add to citation manager EndNote|Ris|BibTeX
| Function | Primer | Sequence (5′-3′) a | Reference |
|---|---|---|---|
| RT-PCR SCE1 primer | SCE1-F | AAAGATCTCATGTCGGGAGGGATCGCACGC (BglII) | |
| SCE1-R | GCGGTCACCTCAAAGCAAAGCAGGGTACTG (BstEII) | ||
| amiRNA primer | I miR-s | agTAAAAATGTGATAACCGTCGGcaggagattcagtttga | |
| II miR-a | tgCCGACGGTTATCACATTTTTActgctgctgctacagcc | ||
| III miR*s | ctCCGACCGTTTTCACATTTTTAttcctgctgctaggctg | ||
| IV miR*a | aaTAAAAATGTGAAAACGGTCGGagagaggcaaaagtgaa | ||
| G-11491 | ccctcagatctCAGCAGCAGCCACAGCAAA (BglII) | Warthmann et al, 2008 | |
| G-11494 | ctcggtcaccGCTGCTGATGCTGATGCCAT (BstEII) | Warthmann et al, 2008 | |
| Hygromycinr gene primer | hpt-F | GATGCCTCCGCTCGAAGTAGCG | |
| hpt-R | GCATCTCCCGCCGTGCAC | ||
| qRT-PCR primer | q-SCE1-F | ACCCAAGTGCAAGTTCCCACAG | |
| q-SCE1-R | GGATAGTGGCTGGAGACCTGCTAT | ||
| q-Actin-F | AAGGCCAATCGTGAGAAGATGACC | ||
| q-Actin-R | CCACAGGTATTGTGTTGGACTCTG |
Table 1 Primers used in this study.
| Function | Primer | Sequence (5′-3′) a | Reference |
|---|---|---|---|
| RT-PCR SCE1 primer | SCE1-F | AAAGATCTCATGTCGGGAGGGATCGCACGC (BglII) | |
| SCE1-R | GCGGTCACCTCAAAGCAAAGCAGGGTACTG (BstEII) | ||
| amiRNA primer | I miR-s | agTAAAAATGTGATAACCGTCGGcaggagattcagtttga | |
| II miR-a | tgCCGACGGTTATCACATTTTTActgctgctgctacagcc | ||
| III miR*s | ctCCGACCGTTTTCACATTTTTAttcctgctgctaggctg | ||
| IV miR*a | aaTAAAAATGTGAAAACGGTCGGagagaggcaaaagtgaa | ||
| G-11491 | ccctcagatctCAGCAGCAGCCACAGCAAA (BglII) | Warthmann et al, 2008 | |
| G-11494 | ctcggtcaccGCTGCTGATGCTGATGCCAT (BstEII) | Warthmann et al, 2008 | |
| Hygromycinr gene primer | hpt-F | GATGCCTCCGCTCGAAGTAGCG | |
| hpt-R | GCATCTCCCGCCGTGCAC | ||
| qRT-PCR primer | q-SCE1-F | ACCCAAGTGCAAGTTCCCACAG | |
| q-SCE1-R | GGATAGTGGCTGGAGACCTGCTAT | ||
| q-Actin-F | AAGGCCAATCGTGAGAAGATGACC | ||
| q-Actin-R | CCACAGGTATTGTGTTGGACTCTG |
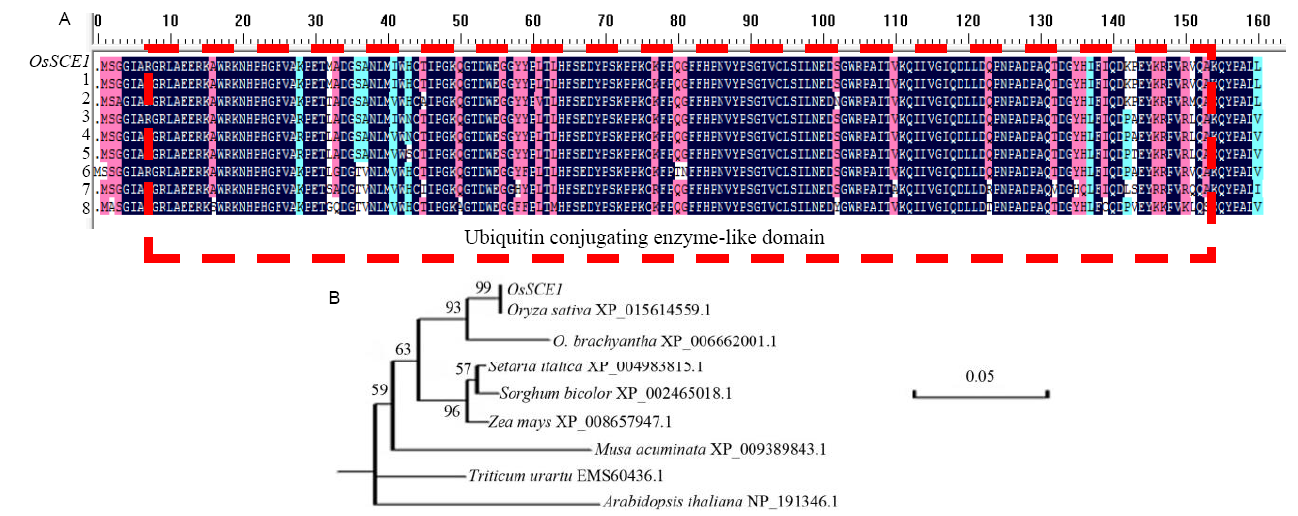
Fig. 1. Multiple sequence alignment (A) and phylogenetic tree analysis (B) of SCE1 protein isolated in this study against reported SCE sequences in database. 1, Oryza sativa XP_015614559.1; 2, O. brachyantha XP_006662001.1; 3, Setaria italica XP_004983815.1; 4, Sorghum bicolor XP_002465018.1; 5, Zea mays XP_008657947.1; 6, Triticum urartu EMS60436.1; 7, Musa acuminata XP_009389843.1; 8, Arabidopsis thaliana NP_191346.1. The colour of alignment indicates conservation among sequences. Dark blue, completely conserved; Pink, ≥ 80% conserved; Cyan, ≥ 50% conserved. Ubiquitin conjugating enzyme-like domains are indicated by the red dashed line. The phylogenetic tree was constructed using the neighbour-joining method (DnaMan1). The numbers at the nodes indicate bootstap values based on 1000 bootstrap replications.
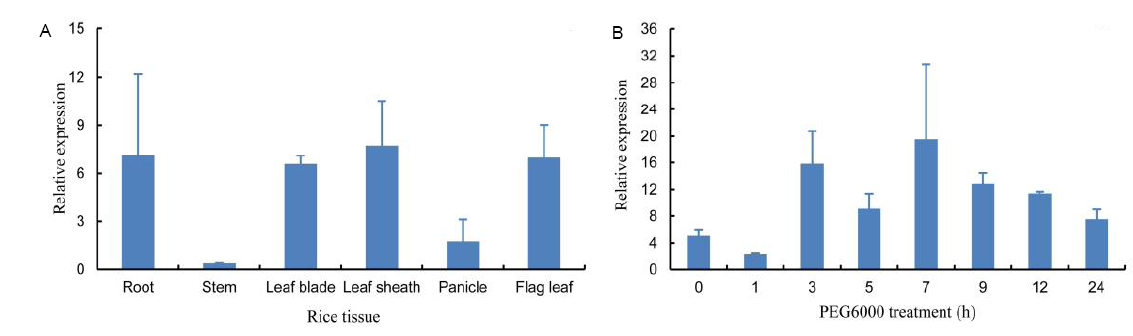
Fig. 2. Expression analysis of OsSCE1 gene. A, Expression pattern of OsSCE1 gene in different tissues of Nipponbare. Data represent Mean ± SD (n = 2). B, Relative expression level of OsSCE1 gene in the leaves of plants treated with PEG6000. Data represent Mean ± SD (n = 3).
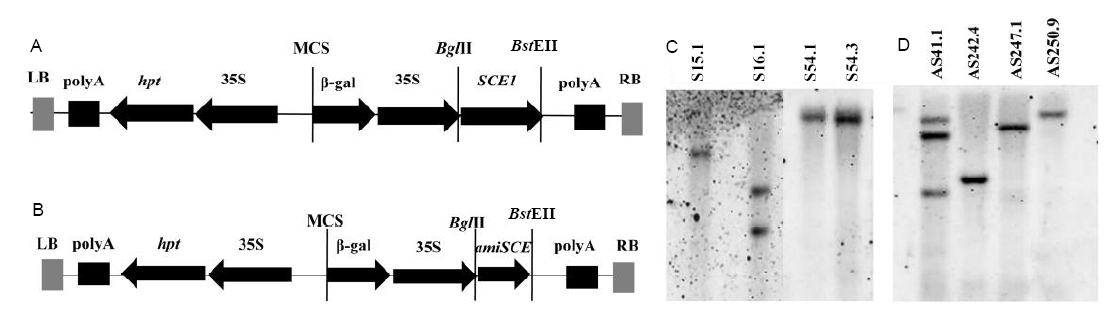
Fig. 3. Overexpression and knock-down analysis of OsSCE1 gene. A, Diagram of P35S:SCE1 construct; B, Diagram of P35S:amiSCE1 construct; C, Southern blot analysis of OsSCE1-OX transgenic plants using hygromycin resistance gene as probe; D, Southern blot analysis of OsSCE1-KD transgenic plants using hygromycin resistance gene as probe.
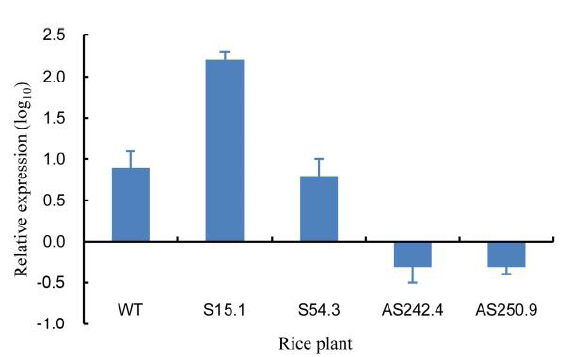
Fig. 4. Relative expression level of OsSCE1 gene in OsSCE1-OX (S15.1 and S54.3) and OsSCE1-KD (AS242.4 and AS250.9) plants under normal conditions. Data represent Mean ± SD (n = 3). WT, Wild type.
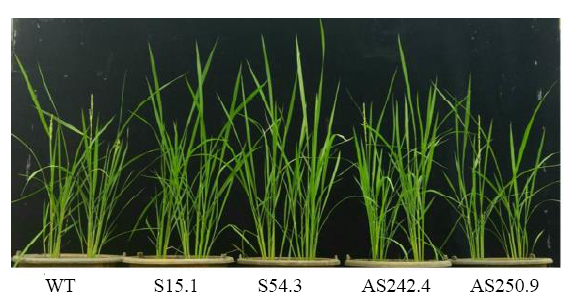

Fig. 5. Morphology of positive transgenic rice lines harboring overexpression (S15.1 and S54.3) and knockdown (AS242.4 and AS250.9) of OsSCE1 gene constructs in comparison with wild type (WT).

Fig. 6. Drought stress tolerance of OsSCE1 overexpressed and knockdown plants. OX1, S15.1; OX2, S54.3; WT, Wild type; KD1, AS242.4; KD2, AS250.9. A, Drought stress tolerance assay of the OsSCE1 transgenic and WT plants under 20% PEG6000 treatment; B, Survival rate of transgenic and WT plants after 7 d of drought stress. Values are survival percentage of plants after PEG6000 treatment (n = 14). Different letters above the bars indicate significant difference at the 0.05 level.
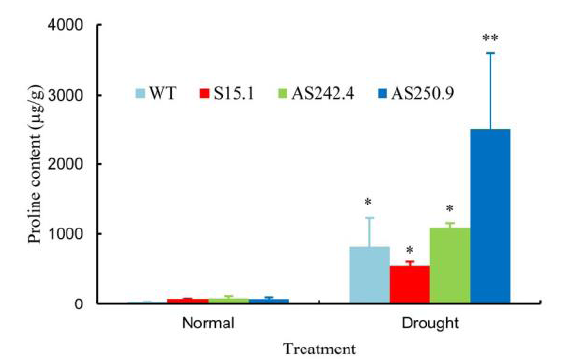
Fig. 7. Proline content in the leaves of 3-week-old transgenic and control plants with or without drought treatment. Values are Mean ± SD of three independent experiments.* and ** represent significant differences at the 0.05 and 0.01 levels, respectively.WT, Wild type.
| [1] | Bates L S, Waldren R P, Teare I D.1973. Rapid determination of free proline for water-stress studies.Plant Soil, 39(1): 205-207. |
| [2] | Castro P H, Tavares R M, Bejarano E R, Azevedo H.2012. SUMO, a heavyweight player in plant abiotic stress responses.Cel Mol Life Sci, 69: 3269-3283. |
| [3] | Castro P H, Couto D, Freitas S, Verde N, Macho A P, Huguet S, Botella M A, Ruiz-Albert J, Tavares R M, Bejarano E R, Azevedo H.2016. SUMO proteases ULP1c and ULP1d are required for development and osmotic stress responses in Arabidopsis thaliana. Plant Mol Biol, 92(1/2): 143-159. |
| [4] | Catala R, Ouyang J, Abreu I A, Hu Y, Seo H, Zhang X, Chua N H.2007. The Arabidopsis E3 SUMO ligase SIZ1 regulates plant growth and drought responses. Plant Cell, 19(9): 2952-2966. |
| [5] | Chaikam V, Karlson D T.2010. Response and transcriptional regulation of rice SUMOylation system during development and stress conditions.BMB Rep, 42: 103-109. |
| [6] | Feng D F, Doolittle R F.1987. Progressive sequence alignment as a prerequisite to correct phylogenetic trees.J Mol Evol, 25(4): 351-360. |
| [7] | Hayat S, Hayat Q, Alyemeni M N, Wani A S, Pichtel J, Ahmad A.2012. Role of proline under changing environments: A review.Plant Signal Behav, 7: 1456-1466. |
| [8] | Hiei Y, Ohta S, Komari T, Kumashiro T.1994. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J, 6(2): 271-282. |
| [9] | Jiang Y J, Qiu Y P, Hu Y R, Yu D Q.2016. Heterologous expression of AtWRKY57 confers drought tolerance in Oryza sativa. Front Plant Sci, 7: 145. |
| [10] | Karan R, Subudhi P K.2012. A stress inducible SUMO conjugating enzyme gene (SaSce9) from a grass halophyte Spartina alterniflora enhances salinity and drought stress tolerance in Arabidopsis. BMC Plant Biol, 12(1): 187. |
| [11] | Kurepa J, Walker J M, Smalle J, Gosink M M, Davis S J, Durham T L, Sung D Y, Vierstra R D.2003. The small ubiquitin-like modifier (SUMO) protein modification system in Arabidopsis. J Biol Chem, 278: 6862-6872. |
| [12] | Lee J, Nam J, Park H C, Na G, Miura K, Jin J B, Yoo C Y, Baek D, Kim D H, Jeong J C, Kim D, Lee S Y, Salt D E, Mengiste T, Gong Q, Ma S, Bohnert H J, Kwak S S, Bressan R A, Hasegawa P M, Yun D J.2006. Salycilic acid-mediated innate immunity in Arabidopsis is regulated by SIZ1 SUMO E3 ligase. Plant J, 49(1): 79-90. |
| [13] | Livak K J, Schmittgen T D.2001. Analaysis of relative gene expression data using real time quantitative PCR and the 2-∆∆CT method.Methods, 25: 402-408. |
| [14] | Lois L M, Lima C D, Chua N H.2003. Small ubiquitin-like modifier modulates abscisic acid signaling in Arabidopsis. Plant Cell, 15(6): 1347-1359. |
| [15] | Miura K, Rus A, Sharkhuu A, Yokoi S, Karthikeyan A S, Raghothama K G, Baek D, Koo Y D, Jin J B, Bressan R A, Yun D J, Hasegawa P M.2005. The Arabidopsis SUMO E3 ligase SIZ1 controls phosphate deficiency responses. Proc Natl Acad Sci USA, 102: 7760-7765. |
| [16] | Miura K, Jin J B, Hasegawa P M.2007a. SUMOylation, a post-translational regulatory process in plants.Curr Opin Plant Biol, 10(5): 495-502. |
| [17] | Miura K, Jin J B, Lee J, Yoo C Y, Stirm V, Miura T, Ashworth E N, Bressan R A, Yun D J, Hasegawa P M.2007b. SIZ1-mediated SUMOylation of ICE1 controls CBF3/DREB1A expression and freezing tolerance in Arabidopsis. Plant Cell, 19(4): 1403-1414. |
| [18] | Miura K, Okamoto H, Okuma E, Shiba H, Kamada H, Hasegawa P M, Murata Y.2013. SIZ1 deficiency causes reduced stomatal aperture and enhanced drought tolerance via controlling salycilic acid-induced accumulation of reactive oxygen species in Arabidopsis. Plant J, 73(1): 91-104. |
| [19] | Nigam N, Singh A, Sahi C, Chandramouli A, Grover A.2008. SUMO-conjugating enzyme (Sce) and FK506-binding protein (FKBP) encoding rice (Oryza sativa L.) genes: Genome-wide analysis, expression studies and evidence for their involvement in abiotic stress response. Mol Genet Genom, 279(4): 371-383. |
| [20] | Park H J, Kim W Y, Park H C, Lee S Y, Bohnert H J, Yun D J.2011. SUMO and SUMOylation in plants.Mol Cells, 32(4): 305-316. |
| [21] | Raorane M L, Mutte S K, Varadarajan A R, Pabuayon I M, Kohli A.2013. Protein SUMOylation and plant abiotic stress signaling: In silico study of rice RLKs, heat shock and Ca(2+)-binding proteins. Plant Cell Rep, 32(7): 1053-1065. |
| [22] | Roberts C S, Rajagopal S, Smith L A, Nguyen T A, Yang W, Nugroho S, Ravi K S, Cao M L, Vijayachandra K, Patell V, Harcourt R L, Dransfield L, Desamero N, Slamet I, Keese P, Killian A, Jefferson R A.1997. A comprehensive set of modular vectors for advanced manipulations and efficient transformation of plants by both Agrobacterium and direct DNA uptake methods. In: Proceedings of Rockefeller Foundation Meeting of the International Program on Rice Biotechnology. 15-20 September. Malacca, Malaysia. |
| [23] | Thompson J D, Higgins D G, Gibson T J.1994. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice.Nucl Acids Res, 22: 4673-4680. |
| [24] | Tomanov K, Zeschmann A, Hermkes R, Eifler K, Ziba I, Grieco M, Novatchkova M, Hofmann K, Hesse H, Bachmair A.2014. Arabidopsis PIAL1 and 2 promote SUMO chain formation as E4-type SUMO ligases and are involved in stress responses and sulfur metabolism. Plant Cell, 26(11): 4547-4560. |
| [25] | Verger A, Perdomo J, Crossley M.2003. Modification with SUMO: A role in transcriptional regulation.EMBO Rep, 4(2): 137-142. |
| [26] | Warthmann N, Chen H, Ossowski S, Weigel D, Hervé P.2008. Highly specific gene silencing by artificial miRNAs in rice.PLoS One, 3(3): e1829. |
| [27] | Xiong H Y, Li J J, Liu P L, Duan J Z, Zhao Y, Guo X, Li Y, Zhang H L, Ali J, Li Z C.2014. Overexpression of OsMYB48-1, a novel MYB-related transcription factor, enhances drought and salinity tolerance in rice. PLoS One, 9(3): e92913. |
| [28] | Xiong R Y, Wang A M.2013. SCE1, the SUMO-conjugating enzyme in plants that interacts with NIb, the RNA-dependent RNA polymerase ofTurnip mosaic virus, is required for viral infection. J Virol, 87(8): 4704-4715. |
| [29] | Zhang S C, Qi Y L, Liu M, Yang C W.2013. SUMO E3 ligase AtMMS21 regulates drought tolerance in Arabidopsis thaliana. J Integr Plant Biol, 55(1): 83-95. |
| (Managing Editor: Li Guan) |
| [1] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [2] | Md. Dhin Islam, Adam H. Price, Paul D. Hallett. Effects of Root Growth of Deep and Shallow Rooting Rice Cultivars in Compacted Paddy Soils on Subsequent Rice Growth [J]. Rice Science, 2023, 30(5): 459-472. |
| [3] | Sheikh Faruk Ahmed, Hayat Ullah, May Zun Aung, Rujira Tisarum, Suriyan Cha-Um, Avishek Datta. Iron Toxicity Tolerance of Rice Genotypes in Relation to Growth, Yield and Physiochemical Characters [J]. Rice Science, 2023, 30(4): 321-334. |
| [4] | Blaise Pascal Muvunyi, Lu Xiang, Zhan Junhui, He Sang, Ye Guoyou. Identification of Potential Zinc Deficiency Responsive Genes and Regulatory Pathways in Rice by Weighted Gene Co-expression Network Analysis [J]. Rice Science, 2022, 29(6): 545-558. |
| [5] | Yousef Alhaj Hamoud, Hiba Shaghaleh, Wang Ruke, Willy Franz Gouertoumbo, Amar Ali Adam hamad, Mohamed Salah Sheteiwy, Wang Zhenchang, Guo Xiangping. Wheat Straw Burial Improves Physiological Traits, Yield and Grain Quality of Rice by Regulating Antioxidant System and Nitrogen Assimilation Enzymes under Alternate Wetting and Drying Irrigation [J]. Rice Science, 2022, 29(5): 473-488. |
| [6] | Chen Eryong, Shen Bo. OsABT, a Rice WD40 Domain-Containing Protein, Is Involved in Abiotic Stress Tolerance [J]. Rice Science, 2022, 29(3): 247-256. |
| [7] | Chen Wei, Cai Yicong, Shakeel Ahmad, Wang Yakun, An Ruihu, Tang Shengjia, Guo Naihui, Wei Xiangjin, Tang Shaoqing, Shao Gaoneng, Jiao Guiai, Xie Lihong, Hu Shikai, Sheng Zhonghua, Hu Peisong. NRL3 Interacts with OsK4 to Regulate Heading Date in Rice [J]. Rice Science, 2022, 29(3): 237-246. |
| [8] | Muhammad Mahran Aslam, Muhammad Abdul Rehman Rashid, Mohammad Aquil Siddiqui, Muhammad Tahir Khan, Fozia Farhat, Shafquat Yasmeen, Imtiaz Ahmad Khan, Shameem Raja, Fatima Rasool, Mahboob Ali Sial, Zhao Yan. Recent Insights into Signaling Responses to Cope Drought Stress in Rice [J]. Rice Science, 2022, 29(2): 105-117. |
| [9] | Balija Vishalakshi, Bangale Umakanth, Ponnuvel Senguttuvel, Makarand Barbadikar Kalyani, Prasad Madamshetty Srinivas, Rao Durbha Sanjeeva, Yadla Hari, Madhav Maganti Sheshu. Improvement of Upland Rice Variety by Pyramiding Drought Tolerance QTL with Two Major Blast Resistance Genes for Sustainable Rice Production [J]. Rice Science, 2021, 28(5): 493-500. |
| [10] | Vera Jesus Da Costa Maria, Ramegowda Yamunarani, Ramegowda Venkategowda, N. Karaba Nataraja, M. Sreeman Sheshshayee, Udayakumar Makarla. Combined Drought and Heat Stress in Rice: Responses, Phenotyping and Strategies to Improve Tolerance [J]. Rice Science, 2021, 28(3): 233-242. |
| [11] | Baoxiang Wang, Yan Liu, Yifeng Wang, Jingfang Li, Zhiguang Sun, Ming Chi, Yungao Xing, Bo Xu, Bo Yang, Jian Li, Jinbo Liu, Tingmu Chen, Zhaowei Fang, Baiguan Lu, Dayong Xu, Kazeem Bello Babatunde. OsbZIP72 Is Involved in Transcriptional Gene-Regulation Pathway of Abscisic Acid Signal Transduction by Activating Rice High-Affinity Potassium Transporter OsHKT1;1 [J]. Rice Science, 2021, 28(3): 257-267. |
| [12] | Panda Debabrata, Sakambari Mishra Swati, Kumar Behera Prafulla. Drought Tolerance in Rice: Focus on Recent Mechanisms and Approaches [J]. Rice Science, 2021, 28(2): 119-132. |
| [13] | Shuting Yuan, Chunjue Xu, Wei Yan, Zhenyi Chang, Xingwang Deng, Zhufeng Chen, Jianxin Wu, Xiaoyan Tang. Alternative Splicing of OsRAD1 Defines C-Terminal Domain Essential for Protein Function in Meiosis [J]. Rice Science, 2020, 27(4): 289-301. |
| [14] | Vijayaraghavareddy Preethi, Xinyou Yin, C. Struik Paul, Makarla Udayakumar, Sreeman Sheshshayee. Responses of Lowland, Upland and Aerobic Rice Genotypes to Water Limitation During Different Phases [J]. Rice Science, 2020, 27(4): 345-354. |
| [15] | Hussain Kashif, Yingxing Zhang, Anley Workie, Riaz Aamir, Abbas Adil, Hasanuzzaman Rani Md., Hong Wang, Xihong Shen, Liyong Cao, Shihua Cheng. Association Mapping of Quantitative Trait Loci for Grain Size in Introgression Line Derived from Oryza rufipogon [J]. Rice Science, 2020, 27(3): 246-254. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||